Evidence for Evolution
A collection of evidence that support the theory of evolution.

A collection of interesting evidence to show how biological evolution explains the diversity of life from across as many disciplines as possible.
CONTENTS
- Direct Observation
- Genetics
- Molecular Biology
- Paleontology and Bioanthropology
- Geology
- Biogeography
- Comparative Anatomy
- Comparative Physiology
- Developmental Biology
- Population Genetics
- Metagenomics
- Physics and Chemistry
- Applications of Evolution
1. DIRECT OBSERVATION
Microevolution (adaptation and other changes within a species) is commonly observed, but the more striking consequences of evolution usually take place on timescales far too long to observe from start to finish. However, there are some well-established cases where macroevolution can be observed in real time. Some definitions here will be useful:
- Biological species concept ~ a species is any group who is reproductively isolated from other such groups, due to e.g. behavioural isolation, genetic incompatibility or failure to produce viable offspring. This is the most common species concept for studying extant life, but is undefined for asexual organisms (prokaryotes), so another concept is required.
- Phylogenetic species concept ~ a species is the smallest monophyletic grouping when performing comparative genomic analysis on a population. This is much more suited for prokaryotes, defining species via genetic similarity.
These multiple definitions are necessary since ‘species’ is not a fundamental unit of nature, but rather a human construct to help us understand biodiversity. This ‘fuzziness’ of the boundaries is expected under evolution which predicts continuous change. Many other species concepts also exist, each useful in their own ways.
- Speciation ~ formation of more than one species from a population of one species, where species is defined suitably using one of the species concepts (like the above).
- Macroevolution ~ variations in heritable traits in populations with multiple species over time. Speciation marks the start of macroevolution.
Although microevolution is useful for conceptualising how Darwinian evolution works (e.g. adaptation: heritable changes with natural selection), it is generally not contested by critics of evolutionary theory. Therefore, we will list here only examples of macroevolution that have been observed in real time. Most of these are in the wild, with a few lab-based studies included too.
The argument
Darwin’s theory of evolution was conceived as the logical conclusion from a series of observations he made during his voyage on the HMS Beagle in the 1830s-50s:
- Organisms are well suited (adapted) for their environment.
- Life shares many characteristics (traits) despite rich variation in form and function.
- Populations can only expand to sizes sustainable by the available resources.
- Populations are naturally generally stable in size (resources are the limiting factor).
- Variation between individuals affects access to resources, influencing their reproduction rates.
- Variation is heritable from parent to offspring.
- The environment (and therefore what is beneficial to organisms) is constantly changing.
Part of Darwin’s main thesis was that evolution must be true unless something is stopping it:
- There is variation in organic beings.
- There is a severe struggle for life.
- Therefore, there must be some variations that are useful to surviving that struggle (from 1 and 2).
- There is a strong principle of inheritance (i.e. offspring are likely to resemble their parents).
- Therefore, useful variations will be preserved (from 3 and 4).
This is the basis for evolution by natural selection (Darwinian evolution). Given that none of the three premises (1, 2 and 4) can be questioned by a sane person, a disbelief in evolution is essentially a pro-active belief in an anti-evolutionary “force”, in the sense that something is actively opposing evolution. Since there is no evidence to believe in such a force, even in the total absence of any evidence for evolution, it suggests that evolution is indeed happening.
Of course in reality, there is ample observable evidence for evolution. If macroevolution can be observed, and we know of no means by which the mechanisms of neo-Darwinian evolution (mutation/selection/drift/gene flow) can stop, and we have consilient evidence indicating continuation of the process back through time, and there is no reason to believe any contrary explanations (e.g. intelligent design/creationism), then the methodologically naturalistic, parsimonious, evidence-driven conclusion follows.
Direct observations are not the best evidence of evolution as a whole: they do not lead directly to the most profound conclusion of evolution, which is universal common descent of all life. Direct observation is just one line of inquiry: the other lines of evidence serve to justify and corroborate the extrapolation of those observations through deep time, synthesising the theory of evolution as we know it.
Lizards with placentas
.](/post/evolution-evidence/lizard_viviparity_hu2301c354ee25b1ddf3c689865b476020_1508251_ffc86fff1a9b0c13b52cda7d8cbca2ab.webp)
Reptiles are known for usually giving birth via egg-laying (oviparity), but there is evidence that some snakes and lizards (order Squamata) transitioned to giving live birth (viviparity) independently and recently. A ’transitional form’ between these two modes is ’lecithotrophic viviparity’, where the egg and yolk is retained and held wholly within the mother.
While observing a population of the lizard Zootoca vivipara in the Alps, reproductive isolation was found between these two subgroups, and attempts at producing hybrids in the lab led to embryonic malformations. Sometimes, the viviparous group would even give birth to two live young and one egg within the same litter of three. The oviparous group is now confined to the range spanning northern Spain and southern France (the Pyrenees), while the viviparous lizards extend across most of Europe. This represents an example of speciation with complete reproductive isolation, together with the gain of a complex new function (viviparity) to boot.
Sources: (Blackburn, 2006), (Cornetti et al., 2015) and here (video).
Fruit flies feeding on apples
.](/post/evolution-evidence/apple_maggot_fly_hue8d28482ff591f37d49c69670e986388_909515_5a5c59ad9e1510a4778c5e8deecdef13.webp)
The apple maggot fly (Rhagoletis pomonella) usually feeds on the berries of hawthorn trees, and is named after apples only because eastern American/Canadian apple growers in 1864 found its maggots feeding on their trees. Since then, the apple-eating and berry-eating groups have become more distinct.
This is a case of ‘sympatric speciation’: the geographic range of the apple group (north-eastern America) is contained within that of the berry group (temperate biomes globally). There is a barrier between the groups because 1) the two trees flower at different times of the year (apples in summer, hawthorns in autumn/fall) so flies must reproduce asynchronously, and 2) each group only lays its eggs on their respective fruit.
Sources: here (several primary sources within).
The London Underground mosquito
.](/post/evolution-evidence/london_underground_mosquito_hu2cf80dceaca8b5e6f047f5d87bbd7801_381541_47de307ab3f58ea606154ff1820e42ca.webp)
They were named due to people being bit by them while hiding in the underground tunnels of London’s tube train network during the Blitz of World War 2. It’s recently been shown that they did not first evolve there. It turns out that the ancestral species, Culex pipiens, lived above ground, while the new species, C. p. f. molestus, evolved in the Middle East ~2000 years ago, adapted to warm and dark below-ground city environments, of which the sealed tunnels of the 1860s London Underground was one.
The new species breeds all-year-round, is cold intolerant and bites rats, mice and humans, while the prior species hibernates in winter. This is a case of ‘allopatric speciation’ (geographic isolation) by ‘disruptive selection’, a rarer type of natural selection where an intermediate trait is selected against while extreme traits are favoured, leading to rapid separation into a bimodal distribution of the two lifecycles. Cross-breeding the two forms in the lab led to infertile eggs, implying reproductive isolation.
Sources: (Garras & Gray, 2022) and (Simms, 2025).
Green algae with multicellularity
.](/post/evolution-evidence/green_algae_multicellular_hu2476658086b3933203d9cc5a8424b5d3_286455_baf9eb50f803f8fcd9291f3b1b0018ce.webp)
‘Colonialism’ (simple clumping/aggregation of single-celled organisms) is well-known, and does not count as multicellularity. But if the cells become obligately multicellular (lifecycle uses clonal division by mitosis and remain together, and splitting them apart kills the organism), the groundwork for de novo multicellularity is laid. This was observed in the lab by introducing a population of green algae (Chlamydomonas reinhardtii, a protist) to cultures of another predatory protist, over a period of 1 year (~750 generations). The strong selective pressure to defend against predation led to obligate multicellularity in the algae. This process, featuring increasingly large clusters of cells, is well-reflected in the extant clade Archaeplastida, which includes green algae (single cell protist), a variety of other colonial protists and plants (complex multicellular).
Another common trait of extant multicellular life is differentiated tissue formation due to cell specialisation. This too has been observed, and represents the formation of complex genetic control systems (by negative feedback loops) as studied by evolutionary developmental biology. Volvox is a good example, being within clade Archaeplastida (above) and having two cell types - one for sexual reproduction, one for phototaxis. Genetics also finds that the famous ‘Yamanaka factors’ for cell differentiation (as well as many other key innovations like cell-to-cell signaling, adhesion and the innate immune system) in animals inherit from those in choanoflagellates (the closest-related protists to animals and our likely last unicellular ancestors). So, both protist-to-plant and protist-to-animal transitions look pretty reasonable on this alone.
Sources: (Herron et al., 2019), (Yamashita et al., 2016), (Gao et al., 2024) for cell specialisation, here (video) and here (long video).
Darwin’s finches, revisited 150 years later
 and their relatedness.](/post/evolution-evidence/darwin_finches_huc9f0c74ccc6b0edd3f76f749474efb18_1027661_d5bc65dbafe1fe4643eeb743ec5d8470.webp)
One of the textbook examples of microevolution is on Darwin’s finches seen during Darwin’s 1830s voyage of the Galápagos islands, where beak size and shape were observed to change in response to food availability during droughts, later revealed to be due to mutations in the ALX1 gene.
More recently, studies from the 1980s onwards have identified speciation in the ‘Big Bird lineage’ on Daphne Major island. Regional droughts which reduce seed dispersal to the islands, such as those that occurred in 1977 and 2004, as well as arrival of competitors, were found to be drivers of selection for beak stiffness. The new lineage of finches reproduces only with its own.
Sources: (Lamichhaney et al., 2017), (Grant & Grant, 2006) and here (article).
Salamanders, a classic ring species
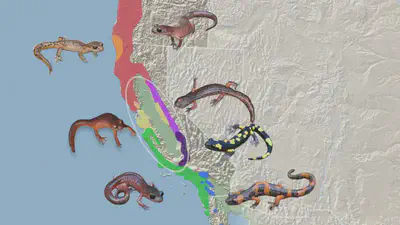
A ‘ring species’ is a rare and aesthetically-pleasing display of speciation wherein a population living outside a circular barrier (e.g. the sands surrounding a lagoon) sequentially mutates and migrates around the circle, so that when they meet up again on the other side, they cannot interbreed. One of the most well-known cases of this is the salamander Ensatina eschscholtzii, which spread around the edge of a dry uninhabitable valley in California. A total of seven subspecies of these salamanders developed around the circle, two of which cannot interbreed with each other. Actually, this case is not a ’true’ ring species, as the diversification process was more complex than simply continuously spreading around the circle, but it still does represent an example of complete speciation.
This process took millions of years, so it wasn’t directly observed, but the studies showing interbreeding capability of neighbouring subspecies despite isolation between two were done in the present, so it’s pretty conclusive as to what happened.
Source: here.
Greenish warbler, another ring species
.](/post/evolution-evidence/greenish_warbler_hu7125a4296bfb02f88d7f225a327d52a2_1340200_24824ba8181b2f3e73982f6584f875e3.webp)
This is another ring species, and one that is closer to a true ring species than the Californian salamanders (though still not a perfect ring species - it seems there are no simple true cases!). These birds, Phylloscopus trochiloides, inhabit the closed boundary of the Tibetan Plateau, of which two reproductively isolated forms co-exist in central Siberia. Genetic studies find some degree of selection against interbreeding, contributing to the speciation process. This happened over about a million years, so we’re using the phylogenetic species concept here.
Sources: here and (Alcaide et al., 2014).
Hybrid plants and polyploidy
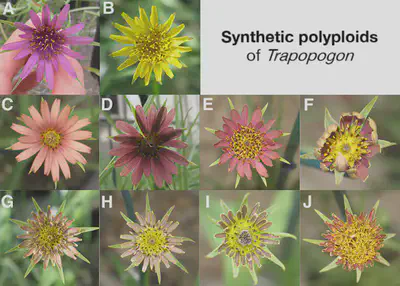

The crossbreeding process that we have used to make new fruits and crops generally exploits polyploidy (e.g. cultivated strawberries) to enhance sensitivity to selection for desired traits. Karpechenko in 1928 was the first to apply this by crossing cabbage (Brassica oleracea) and radish (Raphanus sativus) (both diploids with n = 9). The vast majority of the hybrid seeds failed to produce fertile plants, but a few were fertile and produced highly vigorous offspring with doubled chromosome number (n = 18), and were infertile when crossed with the parent (reproductive isolation). This process also occurs commonly in nature and often results in speciation.
Two new plant species (T. mirus and T. miscellus (goatsbeards)), which are ‘allopolyploid’ plants, have evolved from a common Tragopogon ancestor in the Idaho/Washington region within the last 50 to 60 years, following the introduction of three diploid species from Europe to the US. T. miscellus has become established in the wild and reproduces on its own.
Epilobium angustifolium (fireweeds) has undergone speciation by doubling of the chromosome count, and these autopolyploids cannot produce offspring with the original stock.
Source: (Roose & Gotlieb, 1976), (Soltis et al., 2004), (Mosquin, 1967) and here (info on polyploid speciation)
Amoeboid rhizarian, with endosymbiosis
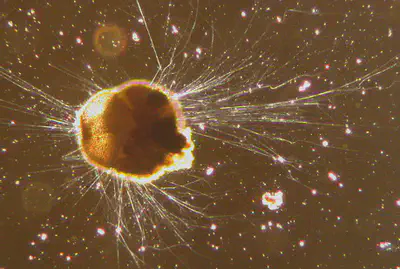
There is excessive evidence that the organelles like mitochondria and chloroplasts (and more recently discovered, the nitroplast) found within extant eukaryotes were originally free-living prokaryotes, which became incorporated (endosymbiosis), but no such thing had been observed, until now. The bacterial order Legionellales are responsible for Legionnaire’s disease and live in water, but are uniquely able to survive and reproduce even after being “eaten” by some amoebae before returning to free-living conditions.
In the lab, it was found that some strains of wild amoeboid protists in clade Rhizaria, class Thecofilosea, were transmitting fully-incorporated Legionellales vertically by cell division. Extracellular transmission of bacteria was not observed, indicating mutualistic endosymbiosis, and genetic studies confirmed divergence of the endosymbiont via a shrinkage of its genome (as expected) and gene translocation to the protist’s nuclear DNA.
Sources: (Solbach, Bonkowski & Dumack, 2021) and (Al-Quadan & Abukwaik, 2022).
Marbled crayfish, with parthenogenesis
.](/post/evolution-evidence/marbled_crayfish_hu6d4848c722310f2b1b87dfda4bc46d54_483656_3cdd3bb134e9a6641ec18f5621905028.webp)
The marbled crayfish (Procambarus virginalis), also known as ‘marmorkrebs’, is an invasive freshwater crayfish species which was first recorded in Germany in the 1990s. Genetic studies trace the whole species back to a single female of the slough crayfish (Procambarus fallax) fished from the Everglades (swamp in Florida) and native to the southeastern US. The marbled crayfish originates entirely from the tropical pet trade, with no known native populations in existence. Since its first discovery, the species has rapidly spread into the wild and is now found abundantly across Europe and in Japan. It is the only species of crayfish reproducing by obligate parthenogenesis (self-reproduction), with only one other species (the red swamp crayfish, Procambrus clarkii) using parthenogenesis only under certain conditions. The marbled crayfish also has a fully triploid genome, while other species are normally diploid, allowing greater variational robustness to environmental change: it can live in acidic, alkaline, polluted and clean water regardless.
Sources: (Gutekunst et al., 2018), here (article), (Scholtz et al., 2003), here (government document) and here (company document).
Cichlid fish
).](/post/evolution-evidence/cichlid_fish_hu5df596704a796cef6d20a234314f2364_551713_24cfbce77eaf1e4af931e49bb63a6729.webp)
Lake Victoria is Africa’s largest lake and a major source for the White Nile river. Lake Victoria’s basin formed 400 kYA, but has been dessiccated (completely dried) three times since then, with the modern Lake Victoria last filling up with water only 15 kYA. Lake Victoria today contains ~500 species of fish, mostly from the subfamily Haplochromini (haplochromine cichlids). Genomic analysis finds that these fish are derived from the cichlids in the geologically older Lake Kivu, currently located ~200 km away, and that three populations of swamp-dwelling cichlids, living in the dry period prior to the last dessication, colonised Lake Victoria and underwent rapid adaptive radiation into the extant biodiversity via repeated hybridisation and specialisation.
Lakes Malawi and Tanganyika also contain the same clade of cichlid fish but are much older, showing steadier rates of evolution over timescales of millions of years. The cichlids have collectively evolved multiple opsin gene variants, with spectral sensitivity tuning correlating with mate choice (colouration for sexual selection) and habitat (deep vs shallow water). Their success and capacity for diversity is strongly attributed to their secondary (pharyngeal) set of jaws, allowing flexible food webs to develop.
Sources: (Verheyen et al., 2003), (Meier et al., 2017), (O’Quinn et al., 2010) on opsins and (Johnson, Kelts & Adada, 2009) on the history of Lake Victoria.
Alligators and chickens growing feathers

In the lab, a change in the expression patterns (controlled by upstream genes) of two regulatory genes led to alligators developing feathers on their skin instead of scales. These occur via the ‘Sonic hedgehog’ (Shh) pathway, one of the many developmental cascades activated by homeotic genes. The phenotypes observed in these experiments closely resembled those of the unusual filamentous appendages found in the fossils of some feathered dinosaurs, as if transitional.
A similar thing has been done to turn the chickens’ scales on their feet into feathers, this time with only one change to the Shh pathway, showing how birds are indeed dinosaurs and descend within Sauropsida.
Sources: (Lowe et al., 2014), (Wu et al., 2017) and (Lee et al., 2020).
Eurasian blackcap
.](/post/evolution-evidence/eurasian_blackcap_hu9a8e6a6672f0b7ad2051aa41a3de533d_944740_a1be45bd2045cf9d811fd1d37cc9e69b.webp)
The migratory bird Sylvia atricapilla typically flies either south-westerly towards Spain or south-easterly into Asia as winter approaches in Europe, but the rise of birdwatching as a hobby in the UK in the 1960s led to a new food source that the westerly-flying birds could migrate to. This change is known to be genetic in basis, involving the magnetoreception-based compass and the circadian rhythym as hybrids fly in intermediate directions. The birds that instead migrated to the British Isles in winter returned home 10 days earlier (due to the shorter distance to central Europe) than those that went towards Spain, and therefore would mate only with themselves (sympatric speciation). The UK-migrating group now has rounder wings and narrower, longer beaks, over just ~30 generations, and although genetic differentiation has not yet reached the point of preventing interbreeding entirely, these birds are quite clearly well on their way to speciation (an incipient species).
Sources: (Torrice, 2009), here (an interview about (Berthold et al., 1992)) and (Mettler et al., 2013).
House mice
In the ~250 years after people introduced house mice to the Faroe Islands, rapid speciation has occured, with noticeable differences in morphology.
Source: (Berry, Jakobson & Peters, 1978).
Worms in the lab
In 1964, six individuals of the polychaete worm Neanthes arenaceodentata were collected from a harbour in southern California and used to found a laboratory population of >1000. By 1991, the lab population could no longer interbreed with the wild population, as there was some premating isolation, and complete postmating isolation. Chromosomal differences were also observed between the lab and wild populations.
Sources: (Weinberg, Starczak and Jörg, 1992) and here.
Fish in Lake Taal
Lake Taal is a lake in the Philippines that was once a bay connected to the South China Sea. In 1754, a volcano erupted and the lake was formed, trapping dozens of saltwater fish. This led to most of their extirpation, but resulted in at least 4 new species, the Freshwater Sardinella (Sardinella tawilis), two gobies Exyrias volcanus and Rhinogobius flavoventris, and the Lake Taal Snake (Hydrophis semperi). It also has a population of Giant Trevally (Caranx ignobilis) that lives in freshwater, compared to its normal saltwater habitat.
Lice on birds
A population of slender pigeon lice (Columbicola columbae), which naturally parasitise rock pigeons (Columba livia), were captured. The lice hide between the parallel feather barbs of the birds to avoid being eaten, requiring them to be below a certain width. The lice were then transferred to giant runts (a domesticated breed of pigeon), which are three times larger, and observed over a period of 4 years (~60 lice generations). Louse body length, metathorax width, and head width were measured, and it was found that the new lice had grown significantly larger. Directional selection is for larger lice on larger hosts, since lice mobility and reproductive capacity is higher without compromising ability to hide within feathers. Partial reproductive isolation was also observed between the two lice groups.
Source: (Villa et al., 2019).
Transmissible cancers and immortalised cell lines
Cancers are clonal populations of cells that can be considered as a ‘breakaway population’ living within a host. While all known human cancers are confined to their host, some cancers are known in animals that are transmissible, essentially becoming their own parasitic species.
- Canine transmissible venereal tumour (CTVT): a sexually transmitted cancer in dogs, genetically found to have originated from a single dog about 6,000 years ago.
- Tasmanian devil facial tumor disease (DFTD): two different cancers on the faces of a species of marsupial, transmissible by biting. DFT1 was first observed in 1996 while the second type DFT2 was discovered in 2014. These cancers have caused severe population decline, leading to endangerment of the species.
- Transmissible cancers of mollusks (Schönbichler & Bergthaler, 2023): cancer cells can be released from bivalve hosts and spread through seawater, surviving host-free for days before infecting new individuals.
- HeLa cell line: immortal cells derived from cervical cancer cells taken from Henrietta Lacks in 1951. The cells are highly aneuploid with 82 abnormal chromosomes. The cells are transmissible between humans via tissue grafts, and are parasitic.
- HEK-293 (human embryonic kidney) cell line: an immortalised cell line commonly used as a eukaryotic target for gene transfection in labs.
This type of breakaway transition has been hypothesised as the distant origin of some evolutionary lineages, such as the single-celled Myxosporea class of cnidarians (jellyfish). This has been referred to as the ‘SCANDAL’ hypothesis (speciation by cancer development in animals: (Panchin, Aleochin & Panchin, 2019)), although this idea remains controversial.
Additional examples
Further collections of observed cases of macroevolution is given on TalkOrigins here and here, known for several decades now.
- Drosophila in the lab: (Ghosh & Joshi, 2012) and (Syed et al., 2017).
2. GENETICS
Genetic similarity between organisms is indicative of evolutionary relatedness, since mutation accumulate within lineages and are passed on to offspring. Studying the genomes of extant life therefore informs evolutionary history.
Pseudogenes
29.49% of the human genome is made up of pseudogenes, the vast majority of which are non-functional (either not transcribed or transcribed levels too low for functionality).
GULO (L-gulonolactone oxidase)
GULO is mostly conserved and subject to purifying selection across the animal kingdom, with a similar gene GLDH appearing in other eukaryotes. It encodes for the enzyme synthesising vitamin C (ascorbic acid) from L-gulono-γ-lactone (in turn from glucose). However, in haplorhines (tarsiers, monkeys, apes: including humans), GULO has been broken by a point deletion (frame shift) mutation in exon 10, so we have to source our own vitamin C from our diet (or die from scurvy). The diet of primates is known to be vitamin C-rich (from fruits), and only a small quantity is required to avoid scurvy, so losing GULO conferred no fitness disadvantage and was lost neutrally to genetic drift. GULO has also been lost independently in the bat genus Pteropus, the domestic guinea pig (Cavia porcellus) and possibly the pika (Ochotona princeps), all of which are monophyletic groups and all are broken in different ways (parsimonious).
In (Mansueto & Good, 2024), it is shown that chromosome 8 (containing GULO) underwent inversion in the haplorhine lineage, but that this did not affect the functionality of GULO. When the pseudogene was formed, the mutation rate increased significantly, indicating loss of a selective pressure, such that the Neanderthal GULO differs from the Homo sapiens GULO by four SNPs.
NANOG (homeobox protein).
Source: (Fairbanks & Maughan, 2006).
DDX11L: 6 copies in chimpanzees, 4 copies in gorillas, and 2 copies in macaques.
Endogenous retroviruses (ERVs)
If a retrovirus infects a germline cell (usually a sperm cell progenitor e.g. spermatocyte), then the viral genome will be inserted inside the germline DNA. When the sperm cell multiplies and fertilises an egg, the viral genome can be passed into the offspring. As long as the virus remains in its dormant state, it will not cause any problems and may become permanently fixed in the genome due to genetic drift. ERV sequences become quickly methylated on inheritance and have their LTRs mutated so cannot jump around the genome any more like retrotransposons, becoming fixed in position before speciation occurs. The viral genome is then said to be ’endogenous’ and will appear in all subsequent descendants of the first infected individual.
We can look for traces of these ’endogeneous retroviruses’ (ERVs) in modern genomes. ERVs can be identified by the long terminal repeats (LTRs) at either end of the genome, and the gag, pol (contains the reverse transcriptase, integrase and protease) and env genes for the viral proteins. Since ERVs insert themselves mostly randomly into the genome, if ERVs are found in extant species with exactly the same positions and identities, it can be safely assumed to be inherited from a common ancestor, as the chance of a coincidental separate identical insertion is negligible.
Most (at least 90%: source) ERVs are non-functional, so the common creationist argument of “common design” loses its validity for ERVs. For example, HERV-W is a human ERV appearing at hundreds of loci in the genome, but only one of them (ERVWE1) is functional and encodes the syncytin-1 protein, which has been exapted and now has an essential function in placental development, conserved in all hominoids (source). Syncytin-2 is similarly exapted from a single locus of a different ERV (HERV-FRD), conserved in all primates. HERV-W is found in both humans and chimpanzees, with 211 of them in humans, 208 of them in chimps, of which 205 of are found in identical locations of both genomes (source). The most likely explanation is that the human-chimp common ancestor had the 205 HERV-W insertions that we both have, and then a few more were acquired independently in each lineage after the split.
We can estimate the probability of this shared ERV distribution occurring under a separate ancestry model, by assuming random insertions of ERVs at sites into each genome. We assume a total of $ N = 10,000,000 $ possible insertion sites in both genomes (‘hotspots’), and compute the probability of having at least $ z $ = 205 shared ERVs given $ a = 211 $ in humans and $ b = 208 $ in chimps as:

$$ P(|X \cap Y| = z) = \frac{\binom{N}{z} \binom{N-z}{b-z} \binom{N-b}{a-z}}{\binom{N}{a} \binom{N}{b}}, \ \ \ \ 0 \leq z \leq \min(a, b) $$
since there are $ \binom{N}{z} $ ways of choosing the intersection, $ \binom{N-z}{b-z} $ ways of choosing the rest of the chimp elements, and $ \binom{N-b}{a-z} $ ways of choosing the rest of the human elements (source). Using the binomial coefficient identity $ \binom{N}{z} \binom{N-z}{b-z} = \binom{N}{b} \binom{b}{z} $ (source), we find it corresponds to the probability mass function (PMF) of a hypergeometric distribution:
$$ P(|X \cap Y| = z) = \frac{\binom{b}{z} \binom{N-b}{a-z}}{\binom{N}{a}} \ \Rightarrow \ |X \cap Y| \sim \textrm{Hypergeometric}(N, b, a). $$
(For a different calculation by Stated Clearly using a slightly simpler model, see here and here.)
Substituting in our numbers, we get (using WolframAlpha):
$$ p = \sum_{z=205}^{208} P(|X \cap Y| = z) = 4.59398489… \times 10^{-1032}. $$
As intuited, the odds of getting the observed HERV-W distribution in humans and chimps without common ancestry is ridiculously tiny: about 1 in $ 10^{1031} $. There are about $ 10^{80} $ atoms in the observable universe, so this is about the same chance of randomly picking the same atom in the universe 14 times in a row! And this is just for one type of ERV in one pair of species - many other types of ERVs are known in many different species (mostly mammals), and they can be used to reconstruct phylogenies in the same way as any other section of the genome.
Molecular clocks for the LTRs of ERVs
We can also study the similarity of the sequences themselves of ERVs in different species. The molecular clock hypothesis assumes a roughly uniform mutation rate to genetic sequences over time, so more closely evolutionarily related species should have more similar sequence identity in their ERVs.
A direct comparison of the LTR sequences of ERVs in different animals demonstrates this correlation with clarity:
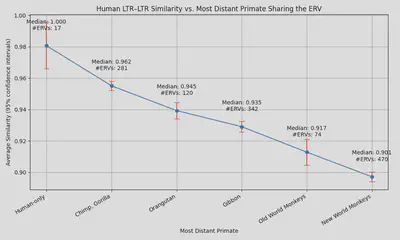
Source: here (reddit post) and here (code and data).
Heat shock proteins
Heat shock proteins (HSPs) are encoded by genes that are turned on in response to high temperature and other stressors. Hsp40 and Hsp70 encode chaperone proteins that ensure reliable protein translation, alleviating the higher risk of misfolding at high temperatures (due to higher molecular kinetic energy). HSPs are therefore critical to the most fundamental functions of life (making proteins) and are therefore expected to be tightly conserved across all life.
As explained in (Gupta & Golding, 1993), HSP70 is in fact the most conserved gene across all life. Furthermore, molecular phylogenetic analysis of HSP70 shows how it demonstrates the origin of eukaryotes from a fusion of an archaeal and bacterial lineage (endosymbiosis), with the eukaryotic HSP70 genes being more closely related to the archaeal and bacterial HSP70 genes respectively than they are to each other.
A more in-depth study was done in (Ravula et al., 2025).
Chromosome 2 fusion in the human lineage
Humans have 23 pairs of chromosomes, while all other great apes have 24 pairs. To explain this discrepancy, it was suggested that two ancestral hominin chromosomes fused to form one in the human lineage. Based on karyotypes, it was predicted in 1972 that human chromosome 2 and chimp chromosomes 12 and 13 (since renamed to 2A and 2B) were homologous and fit the bill for a fusion event. In an end-to-end fusion of two chromatids, we should expect to find:
- a forward and reverse telomere region in the centre (identified by a large number of repeating “TTAGGG” (forward) and “CCCTAA” (reverse) telomere sequences),
- a cryptic (vestigial) second centromere (identified by a long ‘alphoid repeat’ sequence), and
- strong gene homology between the fused chromosome and the two separate chimp chromosomes.
When the genomes were sequenced, it was found that human chromosome 2 indeed satisfied all three of these predictive criteria, demonstrating the fusion beyond all reasonable doubt (Ijdo et al., 1991). The cryptic centromere is degraded and smaller in length (Miga, 2017). The fusion site also contains a particular non-functional pseudogene, DDX11L2, that is only found at telomeres (Costa et al., 2009).
When the Neanderthal and Denisovan genomes were sequenced, they were also found to have the same chromosome 2 fusion as Homo sapiens, indicating the fusion event took place prior to the common ancestor of these species (~750 kYA). Next generation sequencing and molecular clock analyses have estimated that the fusion event occured later than 4.5 MYA, giving a broad range of potential fusion times (Poszewiecka et al., 2022).
Chromosome number alterations are fairly common in nature: for example the many species of horses (genus Equus) vary in chromosome number from 32 pairs to 66 pairs. Fusions are known in extant humans, such as a healthy family with Robertsonian translocations (Song et al., 2016). An evolutionarily recent fusion event is also known in pigs (Thomsen, Høyheim & Christensen, 1996), with a degraded second centromere and homologous banding. The ‘hardest part’ may be getting the fusion to spread (by genetic drift) to fixation in the population rather than the fusion event itself.
Video sources: here (Gutsick Gibbon) and here (Dr Dan).
Beneficial mutations in human evolution
The full genomes of Homo sapiens, Homo neanderthalensis and Denisovans are available, as well as all extant primates, which helps us reconstruct evolutionary relationships and study the origins of individual genes.
Most recent survey of ape genomes: (Yoo et al., 2025).
Human-specific mutations affecting brains and intelligence:
- ARHGAP11: the basal form, ARHGAP11A, encodes the protein RhoGAP with nuclear localisation, found in all extant non-human mammals. A partial duplication ~5 MYA seen in Homo sapiens, Neanderthals and Denisovans led to them additionally acquiring ARHGAP11B, which shows mitochondrial localisation instead. It promotes basal progenitor cells (BP cells) and increases the neocortex size significantly. Sources: here, here and here.
- TKTL1 (transketolase-like 1): modern Homo sapiens has an arginine (R) point mutation (K261R) while Neanderthals, Denisovans, archaic Homo sapiens and other extant primates have the lysine (K) form. Our allele promotes production of basal radial glial cells (bRG cells, neural stem cells), significantly increasing upper-layer cortical neuron production and the size of the brain’s gyri (ridges) in the frontal lobe. Source: here and here (video)
- NOTCH2NL: NOTCH genes prolong proliferation of neuronal progenitor cells and expand cortical neurogenesis. Many of these genes are duplicated in Homo sapiens, Neanderthals and Denisovans to varying degrees. Source: here
- SRGAP2: Partially duplicated to SRGAP2B 3.4 MYA, followed by two larger duplications at 2.4 MYA and 1 MYA. Source: here and here.
- FOXP2: Linked to development of speech and language skills. Source: here.
- TBC1D3: another human-specific gene contributing to the frontal cortex. Source: here
Human-specific mutations affecting muscles and biomechanics:
- PPARGC1A and MYH7: promotes a higher proportion of slow-twitch muscle fibres rather than fast-twitch, favouring endurance and manual dexterity rather than sharp bursts of power. Sources: here and here.
- GDF8 (myostatin): negatively regulates skeletal muscle growth. GDF8 is upregulated (limiting muscle growth) in humans relative to other great apes. Downregulation leads to lower body fat and higher muscle mass (myostatin-related muscle hypertrophy).
- MYH16: changes the musculature of the jaw. Source: here.
- HACSN1: a developmental enhancer leading to limb and digit specialisations. Source: here
Examples from recent human evolution (<300 kYA): source
- ADH1B (alcohol dehydrogenase): the SNP Arg48His is more common in East Asians due to rice domestication, and reduces the risk of alcoholism. Another SNP Arg370Cys occurs in Africa which reduces alcohol dependence.
- PDE10A: leads to enlarged spleens in the Bajau people. The spleen is a reservoir of oxygenated red blood cells, allowing them to hold their breath for longer (hypoxia tolerance) while freediving. Source: here.
- NOS3 (nitric oxide synthase) and others for high-altitude adaptation: in three distinct populations (Tibetans, Andeans and Ethiopians), multiple different mutations in a variety of genes lead to hypoxia tolerance, allowing for their survival at high altitudes.
- Sickle cell trait: in regions of Africa where malaria is prominent, carrying one copy of the recessive sickle cell anaemia allele confers resistance to the Plasmodium parasite. While there are associations of sickle cell trait to other medical conditions, many people with the trait remain healthy, making it net beneficial in malaria-endemic regions. Source: here.
- White skin colour: in northern Europeans, the SLC24A5 gene has an SNP Ala111Thr that leads to decreases melanin expression and hence lighter skin pigmentation, which is beneficial for vitamin D synthesis in the low-sunlight high-latitude regions (where sunburn is less of a risk).
Neutral mutations in recent human evolution
In some of these cases, no harmful effect is observed despite what is typically thought of as a ’loss of function’ (e.g. gene deletion). These result in variation in the population, and may serve as a substrate for future selection, or simply be neutral. Additionally, what is neutral in a current environment may become beneficial in a future environment.
- Blue eyes: leads to blue eyes instead of brown, due to a mutation in OCA2. It has been shown that all blue-eyed people today share a common ancestor living around 6-10 kYA (a perfectly resolved founder event). This is presumed to be a neutral mutation, with the possibility of sexual selection in some cultures. Source: here
- Retention of the median artery into adulthood: normally considered an embryonic structure that regresses around the 8th week of gestation, but it has been found to be retained with increasing frequency in recent times. Source: here.
- ABCC11: the T/T allele carried by nearly all Koreans and many other East Asians is non-functional, preventing its expression. This leads to dry flaky earwax and significantly reduced body odour, even after sweating and exercise. It is so common that deodorant was rarely sold in South Korea until the ~2010s, when cultural and demographic influence created the market. Source: here
- Third molar agenesis: wisdom teeth are becoming less common due to humans eating softer foods that have been processed for ease of consumption, no longer requiring large strong jaws. Associated genes include PAX9, AXIN2, MSX1 and THSD7B. Source: here
Human lactose tolerance
In lactose intolerant people (~65% of humans worldwide), the ability to digest lactose is lost during adolescence. The lactase enzyme is required to metabolise lactose into glucose and galactose. Without lactase in the small intestine, lactose remains available for the bacteria in the large intestine which ferment it, leading to fatty acid and gas production, causing symptoms of lactose intolerance.
The LCT gene codes for lactase, and has a low-affinity promoter. The MCM6 gene, found upstream on chromosome 2, codes for a subunit of helicase (an unrelated protein used in DNA replication), and an intron of MCM6 contains an enhancer for LCT. Transcription factors that bind to the LCT promoter include HNF1-α, GATA and CDX-2, while Oct1 binds to the LCT enhancer.
In mammals, most metabolic genes except lactase are expressed at low levels early in development as nutrients are provided primarily by breast milk, but during adolescence, as these other genes are promoted, low-affinity promoters like LCT are outcompeted, sharply reducing LCT expression. In lactase persistence, point mutations to the LCT enhancer result in an increased affinity for the LCT promoter, allowing it to remain competitive for transcription throughout life, allowing lifelong lactase synthesis. So, this is not a loss of regulation or function, as routinely claimed by ID advocates. Some mutations also reduce the age-related DNA methylation of the enhancer. Lactase persistence has evolved independently with several SNPs (single nucleotide polymorphisms) under strong positive selection in the past 10,000 years of human history, primarily in societies that had dairy farming and pastoralist agriculture.
Sources: here and here (video)
Herbicide and insecticide resistance
https://onlinelibrary.wiley.com/doi/full/10.1111/j.1365-3180.2007.00581.x
https://journals.plos.org/plosgenetics/article?id=10.1371/journal.pgen.0030205
Genomics
The complete genomes of many extant species have been sequenced. Projects in bioinformatics aim to describe and compare the contents and functions of these genomes.
- The Human Genome Project (2003) found that there are about 22,300 protein-coding genes in humans (similar to other mammals) and contains a large number of repetitive sequences (low copy repeats).
- The Chimpanzee Genome Project (2013) revealed that humans and chimpanzees share 99% sequence similarity in protein-coding regions, and 96% sequence similarity across the full genomes, both higher than humans and any other animal.
, [here](https://www.reddit.com/r/DebateEvolution/comments/18uonlm/human_and_chimpanzee_genetic_similarity_an/) and [here](https://www.youtube.com/watch?v=ryBKzJE24Hs).](/post/evolution-evidence/mammal_genome_comparison_huad3c33b20de65bf44ea02984f096a1dc_84920_c4d6f2ef4f1836de17d665d0abb921d9.webp)
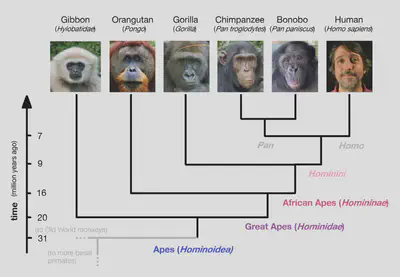
- The ENCODE Project (2021) found that, in humans, about 75-80% of the genome is transcribed into RNAs. However, most of this transcription is either 1) spurious (too low-level to have meaningful biochemical functionality), or 2) forming transposons (LTRs, ERVs, SINEs, LINEs), which simply ‘jump’ around the genome with no useful function. The ENCODE results have been used to imply that “most of the genome is functional”, but ENCODE tested for biochemical activity (of which most is simply low-level noise), not functionality.
Sources: here and (Kellis et al., 2014).
- Among human non-coding DNA, 5% is for gene-related regulatory sequences (promoters, enhancers…). 20% is for introns in genes, most of which serve no function. The rest is completely non-functional (sometimes called ‘junk DNA’, although historically this term was incorrectly applied to all non-coding DNA).

E. Coli citrate metabolism in the LTEE
The Lenski long-term evolution experiment (LTEE) is a famous study that’s been ongoing since 1988, following 12 initially-identical but separate lines of E. coli bacteria over 80,000+ generations thus far. There are no external selective pressures in the LTEE, so the experiment is about what the bacteria could do on their own. Among the outcomes include de novo gene birth from non-coding DNA and near-complete speciation into two variants with differing colony size, but most importantly, one line evolved the ability to eat citrate (Cit) in aerobic conditions, a trait universally absent in wild-type E. coli. This led to an immediate rise in population density.
Contrary to the claims of top intelligent design (ID) proponents (e.g. Dr Michael Behe), this is not merely due to the loss of regulation of CitT (the relevant gene) expression, which would constitute a loss of function. In fact, the CitT gene was in an operon controlled by an anaerobically-active promoter, and underwent gene duplication, and the duplicate was inserted downstream of an aerobically-active promoter. This is therefore a gain of functionality. However, this duplication conferred a negligible (~1%) fitness advantage in the experiment, and at least two other mutations (in an intron of the dctA gene after, and in the gltA gene before) were shown to be necessary to obtain fully-functional citrate metabolism. This therefore meets the criteria for an “irreducibly complex” trait - and it’s one that emerged under experimental conditions normally adverse to innovation (stasis - promotes stabilising selection)!
In an amusing attempt to refute this, ID advocate Scott Minnich (works at Discovery Institute, a politically-motivated creationist organisation) reproduced the experiment in 2016 with a new colony of wild-type E. coli and found the same Cit+ trait emerge! And this time, much faster than in the LTEE, via the same pathway, featuring CitT and dctA. The abstract of their paper ends rather desperately: “We conclude that the rarity of the LTEE mutant was an artifact of the experimental conditions and not a unique evolutionary event. No new genetic information (novel gene function) evolved.” - despite us having disproven that already.
Sources: here, here and here (video).
Tetherin antagonism in HIV groups M, N and O
The human immunodeficiency virus (HIV) is a retrovirus that infects human immune cells expressing the CD4 surface protein, such as helper T-cells and macrophages. Once inside cells, HIV-1’s Nef and Vpu proteins work independently to reduce the expression of CD4, which prevents ‘superinfection’ (two viruses infecting the same cell) and decreases the chance of an immune response. The slow death of helper T-cells leads to a weakened immune system.
If HIV infects a different immune cell, such as a macrophage, the virus’ escape is hampered by the high expression of a cell protein called tetherin. This limits the virulence of HIV in macrophages. However, some strains of HIV-1 have evolved ways to antagonise tetherin using their Vpu and Nef proteins, giving them a second function in addition to retaining their CD4-degrading activity in helper T-cells:
- In HIV-1 group M, tetherin antagonism occurred with 4 concurrent point mutations in Vpu.
- In HIV-1 group N, weak tetherin antagonism occurred with 4 different point mutations in Vpu, but it led to loss of CD4-degrading activity.
- In HIV-1 group O, this occured with just 1 point mutation in Nef (C169S).
So, the same trait evolved two ways (in groups M and O), one of which (group M) was supposedly ‘irreducibly complex’: it was a beneficial trait that required sequential mutations in already functional proteins. Group M now dominates worldwide HIV cases while group O resides mainly in sub-Saharan Africa and group N is very rare.
HIV also simultaneously demonstrates observed ‘macroevolution’ (to the extent that it can be defined for viruses, which are not life). HIV has a zoonotic (animal) origin, as it came from chimpazees’ endemic SIV (simian immunodeficiency virus) strain. SIV is rampant among non-human primates, but each species has evolved to tolerate its own strain. It became human transmissible as HIV in the early 1900s due to mutations that allowed it to bind our CD4 receptors, which differ slightly between humans and other apes, and is far more virulent in humans.
Extra sources: here.
Re-evolution of bacterial flagella
The flagellum is the flagship allegedly irreducibly complex structure, cited ad nauseum by ID advocates. Since it is the one that has been talked about the most, it has also attracted a lot of attention from real scientists who have promptly disarmed it. In one experiment, the master regulator for flagellum synthesis (FleQ) was knocked out, leaving all of the other flagellar genes intact. But under selective pressure for motility, it was found that another transcription factor that regulates nitrogen uptake from the same protein family (NtrC) was able to ‘substitute’ for FleQ, essentially by becoming hyperexpressed, so there’s so much NtrC in the cell that some of it binds to the FleQ-regulated genes and activates them too.
This is an incredibly reliable two-step process, after 24-48 hours we get a mutation in one of the genes upstream of NtrC that leads to higher expression and activation, then within 96 hours of the start we see a second mutation, normally within NtrC itself, that helps fine-tune the expression.
Source: here.
3. MOLECULAR BIOLOGY
rRNA phylogenetics of archaea and eukaryotes
For the earliest stages in evolution (unicellular organisms), fossils are scarce, and genomes have mutated beyond recognition in many places, so we must look more carefully. The ribosome is a key piece of cellular machinery that translates RNA into proteins, whose functionality is so tightly constrained that it can be used to measure relatedness across the whole tree of life.
A surprising result of this analysis is according to ribosome similarities, all eukaryotes descend within archaea. This strongly supports the hypothesis of endosymbiosis, where an ancient archaea cell and an ancient bacteria cell merged to become a eukaryotic cell, with the archaea providing the genes that went into the nucleus.
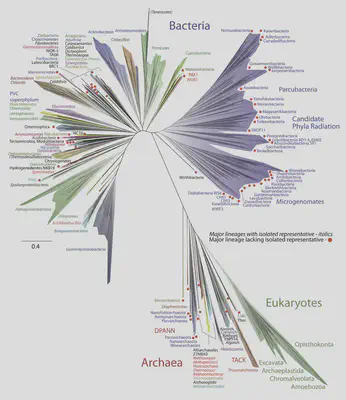
Genomics finds that the most likely matches are from the Asgard clade (discovered in 2012) for the host archaeon, and the phylum Alphaproteobacteria for the endosymbiont. Candidate Asgard clades include Heimdallarchaeota (2019), Wukongarchaeota (2021) and Njordarchaeota (2022). In (Brehmer et al., 2022), it is shown by genomic analysis that mitochondria predates the origin of phagocytosis, so potential endosymbionts may not have been at risk of digestion. In (Imachi et al., 2020), the genome of an Asgardian archaea was found to contain large numbers of homologs for ’eukaryotic signature proteins’ (actins, actin-binding proteins, tubulin, small GTPases, domains for membrane-trafficking proteins), and the morphology of this isolated archaeon had long branched protrusions with no intracellular membranes, similar to the ‘inside-out model’ outlined below.
): explains the origin of the nucleus, mitochondria and the endoplasmic reticulum. **A:** an archaeon (eocyte) and a bacterium living separately. **B:** protrusions (‘blebs’) extended to increase proximity with the bacteria. **C, D:** increasing entanglement. **E:** fusion of internal and external membranes. **F:** fully endogenous with offshoots of internal membranes.](/post/evolution-evidence/endosymbiosis_model_hu2399c2e6c0cb49881036fce3442cd857_373703_8d57c1348242267569e997256c6e143b.webp)
Antibiotic resistance
A striking visualisation of antibiotic resistance is a video of an experiment done by Harvard Medical School, where they created four zones with antibiotic concentrations increasing by a factor of 10 each time. The bacteria would spread outwards up to the boundaries. Most would go extinct, but a few mutants from some populations would survive and move into the next zone. When all zones had been reached, the colonies traced out the pattern of their phylogenetic tree.
. Paper: [here](https://www.science.org/doi/10.1126/science.aag0822).](/post/evolution-evidence/antibiotic_phylogeny_hu61e847926f422580e68087a5a4a947db_1552889_00c3ccfe2c4c88e26cb6fbcaf3658512.webp)
Despite being a highly tangible example of evolution in action, antibiotic resistance is rarely described as “evolution” in the medical literature (sources: here and here).
Herbicide resistance
The first herbicides, 2,4-D (2,4-dichlorophenoxyacetic acid) and 2,4,5-T are synthetic auxins and were discovered due to their selective targeting of dicot plants (includes weeds e.g. dandelions, chickweed, poison ivy etc), while leaving monocot plants (includes crops derived from grasses e.g. wheat, corn, barley) mostly unharmed. The monocots are a monophyletic clade nesting within the angiosperms (flowering plants), indicating an evolutionarily conserved mechanism of molecular tolerance in domestic crops.
Other herbicides like glyphosate target all plants regardless, as they inhibit the more fundamental Shikimate pathway of amino acid synthesis, found in all plants but not in animals. Researchers extracted Salmonella bacteria living in sludge near the glyphosate factories and found them to be resistant due to a mutant EPSP synthase enzyme, and used genetic modification to produce GM seeds confering resistance. Many species of weed have since evolved resistance to glyphosate due to its overuse, with the first occuring as quickly as the GM seeds were invented (Heap & Duke, 2018), and around 60 species resistant as of 2024. This has contributed to the switch in herbicides back to 2,4-D.
(While there is significant controversy surrounding herbicides such as Monsanto’s extremely unethical business practices, industry-funded carcinogenicity studies and the use of Agent Orange, this has no bearing on the underlying biological facts!)
Nylon metabolism in bacteria
Nylon is a synthetic insoluble semi-crystalline polymer of 6-aminohexanoic acid, invented in 1935 and used in a variety of consumer products.
In 1965, Japanese researcher Takashi Fukumura found that 11 bacterial strains in the wastewater of the Toyo Rayon (today Toray) 6-polyamide factory in Nagoya were able to grow on ε-caprolactam, the cyclic amide precursor to Nylon 6. One more species, Corynebacterium aurantiacum, also was found to be able to metabolise linear and cyclic 6-aminohexanoate oligomers. Another group of researchers 4 years later found a strain from the phylum Pseudomonas in the waste water of the same factory that was also able to metabolise 6-aminohexanoate oligomers.
In 1974, Hirosuke Okada conducted research on Flavobacterium also living in the wastewater. He found that the strain Flavobacterium sp. K172 was able to metabolise ε-caprolactam, 6-aminohexanoate and cyclic aminohexanoate-dimer as well as the linear di- bis hexamers of 6-aminohexanoate. The new enzymes had no activity on biologically derived molecules having similar chemical structures.
After some debate in the literature, it has been concluded that one of the new ’nylonase’ enzymes (6-aminohexanoate-dimer hydrolase, EC 3.5.1.46, EII, NylB, P07062) evolved in a two-step process:
- a gene duplication in Flavobacterium to produce a protein named EII’
- base substitutions of EII’ to produce EII.
It has been shown (Negoro et al., 2007) that EII’ has 88% sequence similarity with EII, but only 0.5% of the catalytic activity. Just two point mutations in EII’ found in EII were needed to raise the activity to 85% of EII.
DNT metabolism in bacteria
Dinitrotoluene (DNT) is a unwanted side product in the synthesis of the explosive trinitrotoluene (TNT), which has been produced in the US between 1916 and 1986, and contaminates land surrounding US Air Force bases. Bacteria living in soil polluted with DNT on various bases and DoD sites were found to be consuming the DNT for metabolism as their sole carbon and nitrogen source via aerobic respiration. The DNT was metabolised primarily to nitrite, as well as trace amounts of 2-amino-4-nitrotoluene due to action of nonspecific nitroreductases. These bacteria have been exploited for bioremediation efforts in these areas, with protein engineering helping to increase the degradation rate.
Genetic and molecular studies on one such strain, Burkholderia cepacia R34, found that the bacteria evolved a complex, multiple-protein biochemical pathway by exaptation of proteins with other functions. The novel enzymes included 2,4-DNT dioxygenase (catalyses DNT → 4-methyl-5-nitrocatechol) and methylnitrocatechol monooxygenase (catalyses 4-methyl-5-nitrocatechol → 2-hydroxy-5-methylquinone).
The presence of the transposon remnants and other vestigial genes on the operon strongly suggest the recent evolution of the 2,4-DNT degradation pathway since the extraneous elements have not been eliminated from the region. Comparison with the wild type implies the ancestral natural substrate was naphthalene. Additionally, the transcriptional regulator for the operon does not respond to DNT or its metabolites, instead still recognising salicylate due to its ancestral function. This is a complete metabolic pathway functioning without the regulatory mechanisms that evolve later in more mature pathways, where the regulator mutates to bind a metabolite of the pathway, creating a negative feedback control system.
)](/post/evolution-evidence/dnt_metabolism_pathway_hu18665bab761173a100a2daa46bd4e77c_221616_598f0234bc03c811403f6d1eac6c46ac.webp)
At the Kitzmiller v Dover court case, intelligent design advocate Dr Scott Minnich was presented with this example and he admitted that it satisfied Dr Michael Behe’s definition of irreducible complexity.
Sources: (Kivisaar, 2011), (Pérez-Pantoja et al., 2021), here (document), here (bioremediation), (Leungsakul, Johnson & Wood, 2006) (protein engineering), here and here (TalkOrigins).
De novo promoters and orphan genes
In (Yona, Alm & Gore, 2018), the promoter for the lac operon in E. coli was replaced with random sequences of nucleotides of the same length (~100 nt). A small proportion of these random sequences immediately functioned as low-activity promoters for the operon, but most were inactive. However, after only a single mutation, the whole distribution had shifted towards significantly higher level of lac expression (functionality), with many exceeding that of the wild-type promoter, with not a single one remaining nonfunctional.
Antifreeze proteins
Living in cold environments poses a serious challenge to poikilothermic (not thermally regulated) life, as the water in cells may freeze, halting all metabolic processes and killing the organism. Antifreeze proteins have evolved as a solution: when an ice crystal nucleates, the protein’s ice-binding domain attaches to the surface of the crystal, arresting its growth. The ice-binding domain is a regular arrangement of polar hydrophobic amino acids with a separation very close to the lattice constant of ice, creating an ideal fit for hydrogen bonding and Van der Waals’ forces at the interface.
)](/post/evolution-evidence/antifreeze_protein_mechanism_hua1b4a1aa84e0fbf3cc13876c9df16b1c_329886_87b398b9af5e69e63a1ae1b338e86a5b.webp)
The β-helix motif found in these proteins is common enough in natural secondary structures that it has convergently evolved many times: ice restructing proteins making use of it are known in the winter flounder (Pseudopleuronectes americanus), the spruce budworm (Choristoneura fumiferana), the mealworm beetle (Tenebrio molitor), the snow flea (Hypogastrura harveyi), some sea ice-living diatoms (Fragilariopsis cylindrus) and even some plants like winter rye (Secale cereale). The spruce budworm antifreeze can inhibit freezing as low as -30 °C, below even the metastable supercooling limit of liquid water, but some others only work down to about -5 °C, allowing only marginal additional exploratory capacity in cold environments. These antifreeze proteins have different amino acid composition but all perform the same function. Some protein sequences resemble C-type lectins or sialic acid synthase and have been shown to originate from these by neofunctionalisation (Bogan et al., 2025).
In (Zhuang et al., 2019), it is shown that the antifreeze protein from the northern codfish originated from non-coding DNA, in a process involving frame shift mutations (by 1-nt deletion), duplications and de novo gene birth. Comparison to psueodgenes in closely related species is used to support this. This likely evolved in response to the cyclic northern hemisphere glaciation that began in the late Pliocene (about 3 MYA).
Sources: here, here (video) and here (video).
Cytochrome c oxidase
The cytochrome c oxidase (COX) enzyme is a famous and ubiquitous component of the electron transport chain for respiration, found in bacteria, archaea and the mitochondria of eukaryotes. Since COX is universally conserved, we would expect it to be more similar in closely related organisms, and less so in more distant ones. In fact we find experimentally that there is a strong correlation between the number of amino acid substitutions in the COX enzyme and the time since the divergence of the species. This is a powerful demonstration of the ‘molecular clock’, which gives us an estimate of the time taken for two genomes to have mutated away from a common ancestor, helping us put a time scale onto our evolutionary tree model.
Source: here
Myoglobin
Source: here
Ancient biopolymers
RNA from wooly mammoths, 40 kYA: here (article).
DNA in the environment, 2 MYA: here (paper)
Collagen from dinosaur bone remains (Mary Schweitzer)
Mutations are random
Mutations provide populations with variation, on which the other forces of evolution (selection, drift, gene flow) can act. The Luria-Delbrück experiment (1943) proved that mutations occur randomly with respect to fitness needs (i.e. not directed by the environment, Lamarckian-style). Mutations that appear beneficial in a given environment may have occurred neutrally long before that environment existed, waiting for the right conditions to be selected for.
Although mutations are random, this does not mean that all mutations are equally likely. For example, ’transition’ point mutations (purine to purine, or pyrimidine to pyrimidine) are more common than ’transversion’ point mutations (purine to pyrimidine and vice versa). As a reference, the standard (Watson-Crick) pairing in DNA is:
A (adenine, a purine) binds with T (thymine, a pyrimidine)
G (guanine, a purine) binds with C (cytosine, a pyrimidine)
Epigenetics can also play a role in affecting mutation distributions. For one, mutations are more common in ‘heterochromatin’ (packed DNA, transcriptionally inactive) than euchromatin (loosely packed DNA, transcriptionally active), likely due to reduced access of DNA repair enzymes. Also, since heterochromatin is heavily methylated, methylated cytosines convert to thymine by spontaneous chemical reaction (deamination). The resulting altered distribution of ‘CpG islands’ in the genome can be used to demonstrate common ancestry over intelligent design, as described in this BioLogos article, since it disproves the possibility that genetic differences between clades were chosen for “kind-specific” functionality.
This non-uniform but still random nature of mutations is often described as stochastic, allowing information-theoretic analyses of how evolution impacts genomic diversity (see Section 12).
Natural selection acts on mutations after they occur, often producing predictable patterns that can appear non-random since they have been filtered by survivorship bias. For example, in protein-coding genes, every third nucleotide has a higher chance of a mutation persisting due to the redundancy of the genetic code (synonymous mutations), as quantified by the dN/dS ratio to detect the action of purifying selection on a gene. Meanwhile, in non-functional regions of DNA, mutations occur and fix at the same rate, since no selection filters them out (unconstrained: purely neutral).
The randomness of mutations is fundamentally due to the random nature of quantum mechanics. The nucleobases in DNA undergo spontaneous tautomeric shifts (rapid equilibria) due to the intramolecular quantum tunneling of protons, redistributing the electron density in their aromatic ring systems. This alters the hydrogen bonding environment, so that if the tautomer is present during DNA replication, DNA polymerase may incorporate the wrong complementary base, leading to a point mutation in the complementary strand if not repaired. The mechanism is outlined in detail in Figure 3 of (Tao, Giese & York, 2024).
Other interesting topics in quantum biology include:
- Proton tunnelling in enzyme catalysis: (Klinman, Kohen)
- Proton tunneling in DNA nucleotides: (Slocombe, Sacchi & Al-Khalili)
- Light harvesting in photosynthesis, including Förster resonance energy transfer (FRET) for electron transport between chlorophyll molecules.
- Magnetoreception in birds via radical pair mechanism in cryptochrome proteins.
4. PALEONTOLOGY AND BIOANTHROPOLOGY
NOTE: paleontology is the study of fossils, but since human evolution is a common topic, I include lots of human-specific evidence here from the broader field of bioanthropology, which includes non-fossil evidence.
Fossils are remnants of long-dead life and provide a tangible record of the distant past. We can compare fossilised structures and estimate fossil age using radiometric dating of nearby ash layers to help piece together evolutionary lineages, which can be cross-checked against more precise genetic studies. Taken together, they serve as signposts of how lineages changed over time.
Some of the most obvious evidence for evolution is ’transitional fossils’. Technically, all fossils are ’transitional’, since all life is evolving at all times, but some lineages offer especially clear changes in form over deep time, when ordered by their radiometric dates or strata.
Horse evolution
).](/post/evolution-evidence/horse-fossils_hu09483c634d21f4cef7de49e3c74753b5_953653_ca8b8ea592ee1e7b05098fe63d562859.webp)
Horse feet gradually lost toes, going from 5 toes (small ancestral mammals) to 4 toes on ground (Eohippus) to 4 toes with 3 on ground (Epihippus) to 3 toes (Mesohippus) to 1 toe (Equus). Throughout this process, the carpal bones (homologous with the mammalian wrist) moved upwards due to elongation of the metacarpals, giving the modern appearance of a three-segment leg. The hind limbs also underwent similar later transitions in Mesychippus.
Ancestral horses (Eohippus) originally had toes and lived in more forested areas. The evolution into hooved horses coincided with a changing climate, when the forested environments that Eohippus lived in started to recede and give way to grasslands. While padded feet with toes work fine in forests, they’re not as good as hooves on hard dry soil, especially when you want to move long distances and/or at high speeds.
Over tens of millions of years the smallest digit of the four-toed foot of Hyracotherium becomes drastically reduced leaving its descendants with three toes, and the two lateral toes recede further leaving one big toe with a hoof at the end.
While Eohippus and Hyracotherium had four-toed feet, they could still run on dry grasslands. Horses evolved to become specialised for high bursts of speed and being able to trot or canter for long periods of time in this environment. The evolutionary cost however is that horses are less agile - but life on grasslands doesn’t require much agility since there aren’t many obstacles to manoeuvre around to escape predators.
Sources: here and here (Florida Museum), and here (limbs of the horse)
Bird evolution
One of the most famous transitional fossils known to Darwin, Archaeopteryx displays both ancestral traits (teeth, long bony tail, three claws on wing) and derived traits (feathers, wings, furcula/wishbone, smaller digits).
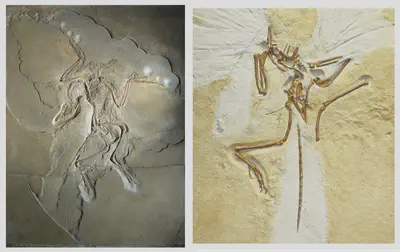
.](/post/evolution-evidence/bird_fossil_hu1ca888a831b451de71ae455244df9aec_2676355_2efff30c2627107db6bafbf3082932b3.webp)
Other points of evidence from the fossil record supporting dinosaur-bird evolution include:
There is a phylogenetically consistent increase in sacral vertebrae number from early archosaurs to theropods to birds, achieved largely by recruitment of adjacent vertebrae into the sacrum via Hox patterning shifts. Early birds retain theropod-like counts, while later birds evolve an expanded, fused synsacrum, supporting a gradual transition rather than a discrete jump.
Early birds retain a theropod-like jugal-quadratojugal arrangement, showing continuity of cheek bone architecture, though progressively modified within clade Avialae.
Early birds retain ancestral theropod skull bones (quadratojugal, postorbital), which are progressively reduced or lost, contributing to the evolution of the highly modified, lightweight avian skull.
Where present, early birds retain the ancestral jugal-postorbital contact seen in archosaurs, supporting skeletal continuity, though this articulation is reduced and eventually lost in modern birds.
Fossils show a progressive reduction of the postorbital and associated bars, which helps decouple skull elements and enables cranial kinesis, a defining feature of modern birds. Developmental data corroborate this via transient embryonic structures.
There is a reduction or loss of the pubic symphysis along the theropod-bird lineage, producing an open pelvis. This reflects broader changes in body plan and soft-tissue systems, not just egg size.
Early birds retain theropod-like cranial pneumatic openings (maxillary/promaxillary fenestrae), which are later reduced or lost as the avian beak (premaxilla) expands.
The key transitional feature is not claws but the gradual fusion and reduction of the hand into a carpometacarpus. Basal birds retain a theropod-like, unfused, grasping hand, demonstrating direct continuity.
The triosseal canal evolved progressively from a more primitive shoulder architecture in theropods, with early birds showing intermediate morphologies that supported limited flight before the fully derived avian flight apparatus emerged.
Genetic data confirm that birds and crocodilians are both archosaurs (nesting within Diapsida, Sauropsida and Tetrapoda), consistent with morphological and fossil evidence placing birds within the dinosaur lineage.
Putative medullary bone in non-avian dinosaurs indicates reproductive physiology similar to birds, supporting deep homology, though some specimens remain debated.
Birds and theropod dinosaurs share a highly similar manual and pedal digit configuration, including reduction from a five-digit ancestor. This reflects a progressive, well-documented pattern of digit loss and specialisation.
Whale evolution
. Many other species with nearly complete fossils are known: e.g. [here](https://en.wikipedia.org/wiki/Evolution_of_cetaceans).](/post/evolution-evidence/whale_fossils_huf29ce38d64c98f8c107428e5f99992df_4415588_d35f17474e7e738a0ff676355af9f252.webp)
. **(c)**: The post-cranial fossil material found from *Indohyus*. **(d)**: Reconstruction of postcrania and artist's impression.](/post/evolution-evidence/indohyus_fossils_hu61c939bece30306a13b349ff092c3b24_2296788_781352e25589f41f4db6d03419d693ec.webp)
Tetrapod evolution
Wikipedia pages: Evolution of tetrapods, Skeletal changes and Vertebrate land invasion.
Plant evolution
Plant fossils has its own field of study: paleobotany.
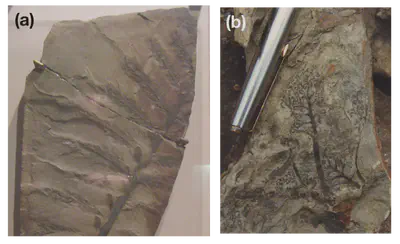

Ediacaran Biota (early animals)
While the ‘Cambrian explosion’ is the most well-known expansion of animal and plant life, responsible for the relatively rapid radiation of animal phyla, it was not the origin of most of these phyla, which appeared in the preceeding period, the Ediacaran (635 - 538.8 MYA).
Geologic regions bearing lots of Ediacaran fossils include the Doushantuo formation in China and the Avalon peninsula in Canada.
The early Ediacaran yields some interesting finds:
- Lantianella (early Ediacaran, 635-590 MYA), likely belonging to Cnidaria.
- Phosphatised animal embryos, such as Megasphaera, Caveasphaera and Helicoforamina.
- Acritarchs: microfossils without an assignment.
However, the later Ediacaran provides most of the better-known biota from this period, divided into the Avalon (571 - 555 MYA), White Sea (560 - 551 MYA) and Nama (555 - 541 MYA) assemblages.
The Avalon biota, following the Avalon explosion, consists of sessile (non-moving) frondose (having fronds) fractal rangeomorphs such as Charnia and Fractofusus (stem non-sponge animals), frond-like arboreomoprhs (having bulb-shaped anchors) like Charniodiscus and disc-like organisms such as Cyclomedusa. Cnidarians include the muscle-bearing Haootia quadriformis.
The White Sea biota includes Tribrachidium (sessile benthic suspension feeder with tri-radial symmetry), Dickinsonia (stem bilaterian), Ikaria (worm-like bilaterian) and Kimberella (stem mollusc or spiralian), representing increasingly modern animal clades.
).](/post/evolution-evidence/white_sea_fossils_hub48a17282c15d6123cf7467f73634eb0_328505_812608a5585b19a1933cfee877e795a2.webp)
The Nama assemblage follows the end-Ediacaran extinction event and coincides with the Baykonurian glaciation. It includes fossils such as Cloudina (tubular and biomineralised, likely an annelid), Yilingia (segmented bilaterian: annelid or panarthropod) and Namacalathus (early relative of brachiopods and bryozoans).
).](/post/evolution-evidence/ediacaran_cambrian_fossils_hu04869a48f430ace6b13551d497a5fa6b_632965_778811570f29315a254abea1ea1e8005.webp)
The Ediacaran biota contain a wide range of genera, and some of these fossils are shown:
 (exceptionally well-preseved soft tissue) and [ichnofossils](https://en.wikipedia.org/wiki/Trace_fossil) (trace fossils).](/post/evolution-evidence/ediacaran_hu419e84fff60dc065fb8d9e8a54a05a09_851791_d941010568a3747e61b499fd2ca916c3.webp)
There are also animal fossils dating to even further back than the Ediacaran:
- The Cryogenian biota, indicated as sponges (animals) by the presence of molecular biomarkers (24-isopropylcholestane, a degraded steroid) in the sediment, dating to around 700 MYA. This use of steranes is discussed in (Love et al., 2009), as well as in (Zumberge et al., 2018) for demosponges.
- Otavia antiqua: a multicellular animal assigned to phylum Porifera (sponges), appearing as far back as 760 MYA (late Tonian). Described in (Brain et al., 2012).
Early Eukaryotes
Two major clades of eukaryotes include:
- Clade Archaeplastida contains red and green algae (multicellular protists), glaucophytes (unicellular protists) and land plants.
- Clade Amorphea contains amoebas, the clade Holomycota (containing fungi) and the clade Holozoa (containing animals).
Some known fossils from some of these clades include:
- Bangiomorpha pubescens: a species of red algae (the complex multicellular clade Archaeplastida) found from 1,200 MYA (late Ectasian and Stenian). The fossil includes differentiated reproductive cells that are the oldest evidence of sexual reproduction. (Butterfield, 2000).
- Horodyskia: a eukaryote dating to the Calymmian, 1,500 MYA.
- The Gaoyuzhuang biota are a group of possible algae (multicellular eukaryotes), such as Grandilingulata, Tuanshanzia, Changchengia, dating to the early Calymmian around 1,590 MYA. These are likely the base of clade Archaeplastida.
- Rafatazmia chitrakootensis and Ramathallus lobatus: crown group red algae dating to the late Statherian, 1,650 MYA.
- Grypania: a tube-shaped fossil proposed to be an algae, dated to the Orosirian, 1,870 MYA.
- Diskagma buttonii: a basal eukaryote of unknown affinity dated to the Rhyacian, 2,200 MYA.
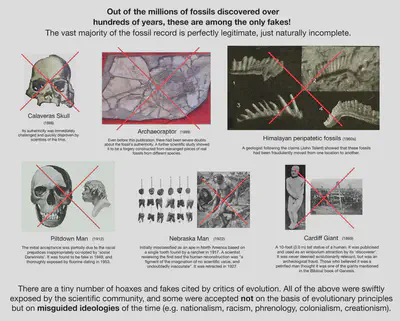
Fake fossils?
The fact that these fraudulent cases are so rare, are so thoroughly well-scrutinised when they do happen, and are always rejected by the scientific community, serves as reassurance that the vast majority of the fossil record is in fact perfectly reliable, just naturally incomplete.
The occurrence of these frauds in the early 1900s led to increased caution and scrutiny on more recent fossil finds, increasing the reliability of the field.
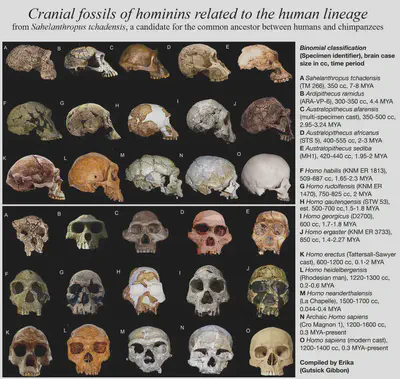
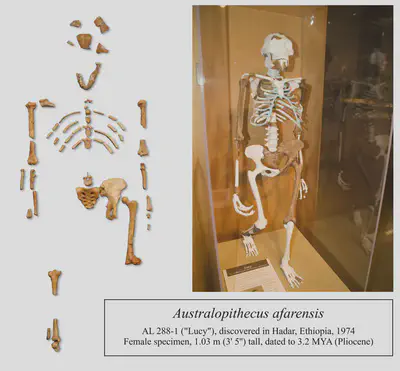
Human evolution
Human evolution is an especially well-studied topic. We are primates and great apes, and there is an abundance of fossils to tell us how our lineages developed over time. Genus Homo arose from the prior genus Australopithecus about 2.5 million years ago, and following a period filled with numerous species of Homo, our species Homo sapiens emerged about 300,000 years ago. Our close relationship with chimpanzees, gorillas and bonobos make them great for studying behaviour, too.
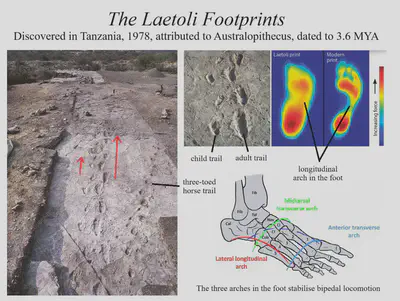




Primate anatomy and taxonomy
The study of extant primates can also give us clues into our shared past.
In 1698, English anatomist Edward Tyson dissected a chimpanzee and noted in his book that the chimpanzee has more in common with humans than with any other ape or monkey, particularly with respect to its brain.
In 1747, taxonomist Carl Linnaeus wrote to J. G. Gmelin, expressing (with circumspect forbearance) his conclusion that humans and other apes must, by the logic of his own nested hierarchies, belong to the same group, which he called Anthropomorpha. He writes:
As a natural historian according to the principles of science, up to the present time I have been not been able to discover any character by which man can be distinguished from the ape; for there are somewhere apes which are less hairy than man, erect in position, going just like him on two feet, and recalling the human species by the use they make of their hands and feet, to such an extent, that the less educated travellers have given them out as a kind of man.
I demand of you, and of the whole world, that you show me a generic character — one that is according to generally accepted principles of classification, by which to distinguish between Man and Ape. I myself most assuredly know of none…. But, if I had called man an ape, or vice versa, I should have fallen under the ire of all the theologians. It may be that as a naturalist I ought to have done so.
These early naturalists (along with many other Western contemporaries) recognised the similarities, but, living prior to Darwin, had no theoretical framework with which to explain them.
). The fingers of gorillas with vitiligo look strikingly similar to those of an adult (White) human's when the dark pigment is removed. The appearance of age in the gorilla fingers relative to humans is due to the autapomorphic [neoteny](https://en.wikipedia.org/wiki/Neoteny_in_humans) driven by [sexual selection](https://en.wikipedia.org/wiki/Sexual_selection_in_humans) in our lineage.](/post/evolution-evidence/gorilla-hands-look-human-pink-pigmentation_hu4212302d0459fb14c8724f574b734d7a_123837_98886c8b4192e4a76e8b60523be6db56.webp)



.](/post/evolution-evidence/barry_63_hu7c2004380a664d8eb7b128867570d792_549157_c1b7030f03e2602b611f4087b6181fb7.webp)
Primate ethology (behaviour)
Primate behaviours are stunningly reminiscent of human behaviours.
Many non-human primates display a clear ’theory of mind’ (the understanding that others’ beliefs, desires, intentions, emotions and thoughts may be different from one’s own). Primatological studies find they:
- warn unaware group members of danger (Crockford et al., 2012)
- preferentially comfort bereaved mother chimps (Goldsborough et al., 2020)
- mediate and resolve fights they were not involved in (de Waal, 2004)
- use tools extensively, such as in nut cracking, termite fishing, spearing bushbabies for hunting etc. (Gibbons, 2007 and Hicks et al., 2019)
- provide a group mate with tools they need for a task they are not involved in (Yamamoto, Humle & Tanaka, 2009)
- use plants containing medicinal compounds to heal serious facial wounds, chewing them up and packing them in (Laumer et al., 2024)
- exhibit reconciliation, consolation and empathy
- mourn their dead (source: here)
- engage in vicious warfare against other groups, torturing rival males to death and assimilating rival females into their group (Wilson & Wrangham, 2003)
- conspire to instigate violent coups against alpha males (Jensen, Call & Tomasello, 2007)
- understand justice and fairness, with a similar level of prosociality as human toddlers (Proctor et al., 2013)
- easily pass the ‘mirror test’ (recognise themself in the mirror and use it to groom themselves - easily observed at zoos)
- rescue nonrelated group members who are in mortal danger e.g. drowning in deep water (Fouts & Mills, 1997)
- perform worse at easier tasks and better at harder tasks when being watched by people than when alone, as if they were feeling the pressure/anxiety to perform in front of an audience (reputation management) (Lin, Muramatsu & Yamamoto, 2024)
- dance for fun and to attract males (Coy, Caspar & Patel-Grosz, 2025)
- point at things to tell group mates who haven’t seen something before, demonstrating theory of mind (Townrow & Krupenye, 2025)
- have ‘circles of friends’ with social grooming (van Leeuwen et al., 2025)
- rationally revise their beliefs based on new evidence (Schleihauf et al., 2025)
Primates also have complex language capabilities:
- Gelada vocalisations follow fundamental linguistic rules of grammar called Zipf’s law and Menzerath’s law (Gustison et al., 2016)
- Campbell’s monkeys and panins both use syntax and grammar (Ouattara, Alban & Klaus, 2009)
- Marmosets in groups call to each other with specific sounds, as if they have names (Oren et al., 2024).
- Chimpanzees and human toddlers have a ~90% overlap in ‘innate’ gestures (Kersken et al., 2019)
- Chimpanzees have unique gestures within families, a necessary precondition for inventing de novo language (van Boekholt et al., 2024)
- Bonobos combine vocalisations in non-trivial ways to create new meanings (Berthet, Surbeck & Townsend, 2025)
Many of these behaviours were at one point (even recently) thought to be the unique characteristic of humans that sets us apart, but in fact they are merely differences in degree rather than kind.
Many primatologists doing fieldwork regularly observe the ‘humanity’ in chimpanzees in particular.
Bipedalism in Hominins
Walking up on two feet (bipedalism) is a trait unique to humans today among the primates, so studying how this evolved is important. The suite of characteristics indicative of bipedalism, which originated in late Miocene hominids, is:
- Anterior foramen magnum *: skull rests on the top of the spine.
- Sagittally-oriented iliac blades (bowl-shaped pelvis) *: pelvis rests upright, supports visceral organs around the abdomen.
- Valgus knee (bicondylar angle) *: femur angled to keep knees in line.
- In-line hallux: big toe is aligned with the other toes, aiding in walking.
- Lumbar lordosis (S-shaped vertebral column): supports upright gait.
- Arched foot: three arches (medial, lateral, transverse) in the feet act as shock absorbers in walking.
* strongest indicators, since these biomechanically prevent quadrupedalism.
Fossils of extinct primates can be analysed to see whether these traits are present, allowing us to trace the gradual evolution of bipedalism. For example:
- Ardipithecus ramidus has a ‘mosaic’ pelvis: the ilium (top) is human-like, while the ischium (bottom) is more ape-like. It has an anterior inferior iliac spine, the muscle attachment site indicative of bipedalism. Its foramen magnum is also intermediate between a human and a chimp.
- All species of Australopithecus have the foramen magnum in the anterior position, bowl-shaped pelvis, valgus knee, inline hallux, smaller back curvature and 12 thoracic vertebra. They had a partial third (transverse) arch in their feet (transitional).
Presentation with extensive comparisons: here by Gutsick Gibbon.
Bones can ‘follow’ any biomechanical trend demanded during evolution, due to Wolff’s law of causal morphogenesis. This allows relatively fast variation in bone shape depending on the behaviour of organisms.
Taxonomy of Australopithecus and Homo habilis
Our genus, Homo emerged from one of the coexisting species of the genus Australopithecus. Based on the available fossil record, this transition appears to be quite subtle: the brain case sizes overlap, both can use stone tools, both have similar dentition (teeth), and biomechanics studies indicate both were mostly bipedal (though A. afarensis only had two arches plus a less-curved third arch in the foot, suggesting habitual bipedality, while H. habilis had three fully formed arches, suggesting full obligate bipedality). This high degree of similarity has even led to some paleoanthropologists to suggest that H. habilis should in fact be Australopithecus habilis. Although this has not been formally adopted, the challenge of a clear-cut taxonomic classification demonstrates the highly transitional nature of these species.
Another example of this comes (ironically) from creationists, who are ideologically required to divide the hominin fossil record into two allegedly mutually exclusive groups: the ‘ape’ kind and the ‘human’ kind. However, six famous hominin cranium fossils of H. habilis and early H. erectus (KNM-ER 1813, Java man, Peking man, KNM-ER 1470, KNM-ER 3733 and Turkana Boy) were all classified by seven different creationists completely differently, precisely as expected of a ’transitional’ species without a true clear divide.
Neanderthals are not our ancestors
One of our closest relatives, the Neanderthals, went extinct about 40,000 years ago. Autapomorphies (uniquely defining traits) of Homo neanderthalensis include retreating cheekbones (zygomatics), the occipital bun, large nasal aperture, enhanced prognathism, enhanced brow ridges (supraorbital torus), platycephalic skull, angled squamosal suture, retromolar gap and an elliptical foramen magnum.
Significant hominin fossils, ichnofossils and artifacts
- Ardi: partial skeleton of Ardipithecus ramidus, 4.4 MYA, discovered in the Afar rift valley (Ethiopia).
- Little Foot (StW 573): near-complete specimen, Australopithecus africanus, 3.67 MYA. Found in the ‘Cradle of Humankind’ in South Africa, an area home to many other early hominins.
- Burtele foot: partial foot, 3.4 MYA, with a divergent hallux, assigned to Australopithecus deyiremeda.
- Taung Child: skull of a 3-year-old Australopithecus africanus, 3.3 MYA.
- Dikika Child (Selam): skull of a 3-year-old Australopithecus afarensis, 3.3 MYA.
- Lucy (AL 288-1): partial skeleton of Australopithecus afarensis, 3.2 MYA.
- Ledi Geraru mandible (LD 350-1): jaw assigned to basal genus Homo, 2.78 MYA.
- Mrs Ples: complete skull of Australopthecus africanus, 2.35 MYA.
- Malapa hominins (MH1 and MH2): two partial skeletons of Australopithecus sediba, 2 MYA.
- Dmanisi skulls: a set of skulls of Homo erectus, 1.81 MYA.
- Turkana Boy: near-complete young Homo erectus, 1.55 MYA.
- Peking Man: a Chinese specimen of Homo erectus, 500 kYA.
- East Asian archaics: a collection of late Homo erectus, archaic Homo sapiens and Denisovan specimens from China and surroundings, e.g. Dali man, Xiahe mandible, Harbin skull/Dragon man…
- Denny: teeth from a 13-year old female containing DNA, Neanderthal-Denisovan hybrid, 90 kYA.
- Laetoli footprints: two trackways, attributed to Australopithecus, 3.6 MYA. The indentations suggest a fully in-line hallux.
- Lomekwi stone tools: oldest stone tools known, predating genus Homo.
Evolution of tool use: paleolithic stone age technology
Archaeological finds associated with hominin remains often include stone tools and other objects of interest. The progression in the complexity of these tools follows evolutionary lines, and is divided into industries:
- Lomekwian tools: from 3.3 MYA, knapped rocks attributed to Australopithecus or Kenyanthropus.
- Oldowan tools: from 2.9 MYA, attributed to Homo habilis and possibly Australopithecus garhi.
- Acheulean tools: from 1.5 MYA, attributed to Homo erectus and Homo heidelbergensis.
- Mousterian tools: from 300 kYA, attributed to Homo neanderthalensis.
- Aurignacian tools: from 50 KYA, attributed to anatomically modern Homo sapiens.
Once Homo sapiens evolved, tool complexity increased rapidly since ~50 kYA, culminating in the advent of civilised societies.
In (Braun et al., 2025), it is found that extant wild chimpanzees choose tools in a similar way to those found in the Oldowan tool industry, selecting harder stones for hammers and softer stones for anvils.
Control of fire for the purposes of cooking meat has been attributed to Homo erectus over 1 MYA, with confidence increasing for more recent use in heat treatment of stone weapons. Use of fire for processing metals did not occur until very recently (the iron/copper/bronze ages of recent civilisation).
5. GEOLOGY
Stratigraphy
The idea that rocks are deposited in layers (strata: older below, younger above) has been known since Steno in the 17th century (the ’law of superposition’). It is therefore usually the case that fossils found in deeper layers are older than those found above, serving as a rough guide to their age (a qualitative, relative dating method). However, other geologic processes like erosion, folding and faulting can disrupt this order occasionally, so more reliable references are needed.
Fossil species that are used to distinguish one layer from another are called index fossils, which occur for a limited interval of time. Usually index fossils are fossil organisms that are common, easily identified, and found across a large area. When a fossil is found, the nearest volcanic ash layers above and below it can be radiometrically dated, allowing us to bound the age of the fossil (tephrochronology).
In (Benton & Hitchen, 1997), the existing fossil record for 384 different clades all across the animal kingdom was surveyed and cross-referenced with their claimed evolutionary relationships. Using three different statistical metrics (Spearman’s rank coefficient, and two others dedicated to quantifying the presence of ‘ghost lineages’), it was found that all three falsify the null hypothesis (if the fossil record does not reflect the major patterns of evolution, there would be no evidence for congruence between the two sets of data in our random sample of cladograms).
Dendrochronology
Trees grow at a rate of approximately 1 ring in their trunk per year. By sampling the rings of multiple trees in a given region, and matching the thicknesses of each ring to the others, we can estimate the age of trees. This also allows for identification of missing or additional rings in a given tree, which are indicative of ecological disturbances (e.g. wildfires, insect outbreaks…).
Dendrochronology can be used to date wooden artifacts in archaeology from about 10000 years ago to present, since the last ring can be matched to the year it was cut down and used. The carbonised wood in charcoal can be both carbon dated and dendrochronologically dated, allowing us to cross-reference the two methods. We can also infer the climate conditions over its lifetime (paleoclimatology), inferring the past temperature, precipitation and cloud cover from the ring data. Climate data can also be cross-referenced against other sources (e.g. ice cores, sediment cores, historical records, meteorological data…). Examining the trace mineral content of the rings (e.g. carbon-12/13 ratio) provides further data on atmospheric conditions (stable isotope dendrochronology).
Source: here
Ice core dating
…
Varve chronology
Sedimentary layers in lakes and oceans.
Source: here
Paleomagnetic dating
, the rates of tectonic plate spreading at mid-oceanic ridges can be measured by recording the magnetisation of the sediment, which tracks the orientation of the geomagnetic field at its formation. This was one of the early 'smoking guns' used to support tectonic theory, and also allows dating by measuring continental drift, helping to calibrate magnetic analyses elsewhere by correlating with geomagnetic field reversals.](/post/evolution-evidence/geomagnetic_striping_hu5b1cf3e542791493dff0ea0d47ab6fe9_685745_7aba922bce26df11d418e8910ca06f2d.webp)
Iron-60 deposits in magnetofossils
Magnetotactic bacteria (MTBs) live a few centimetres below the sediment on an ocean floor. When sediment deposits on the ocean floor, the MTBs move up to maintain their depth. MTBs uniquely contain ferromagnetic particles which they use to passively align themselves to the geomagnetic field (magnetoreception). These particles are produced by the MTBs consuming iron hydroxide. When they die, their filaments retain the iron, forming a column of ‘magnetofossils’ where depth correlates with age.
In 1999, a new isotope of iron was discovered on the deep seafloor, iron-60 ($ ^{60}Fe$), in polymetallic nodules. $ ^{60}Fe$ is radioactive with a half-life of 2.6 million years, and cannot be formed by stellar nucleosynthesis, so its only plausible origin is from a distant supernova showering the Earth with $ ^{60}Fe$. Using beryllium dating and stratigraphy, a sharp increase in the $ ^{60}Fe/Fe$ ratio of MTB magnetofossils was observed between 2.7 - 1.7 MYA.
Calculations were performed to estimate the required position, distance and stellar mass of a potential supernova that could be responsible. A supernova observed to have occurred within the Tuc-Hor stellar group ∼2.8 Myr ago, 330 light years away, with supernova material arriving on Earth ∼2.2 Myr ago, was identified as the likely source. It was found that the iron-60 deposits are consistent with turbulent radioisotopic transport in dust grains originating from this supernova explosion.
Sources: (Ludwig et al., 2016), (Fields et al., 2019) and here (article).
Oklo natural nuclear reactor
In 1972, an anomaly in uranium isotopes was found at a mining site in Oklo, Gabon (Africa), with suspicions of secret nuclear enrichment by a rogue state. Subsequent analysis however found that isotopic data from other metals yielded the conclusion that nuclear fission had been occurring at this site around 2 billion years ago, naturally reducing the abundance of the fissile $ ^{235}$U isotope. Other sites with similar activity have also been found nearby (Gauthier-Lafaye, Holliger & Blanc, 1996).
The data from Oklo has also been used to check that the fine structure constant ($ \alpha = 0.007297… \approx 1/137 $) has remained truly constant over deep time. $ \alpha $ is the dimensionless parameter in relativistic quantum theory and therefore governs electronic interactions in radioactivity. Cosmological observations also verify this fact with even better confidence, validating geophysical uniformitarianism (Davis & Hamden, 2015).
Extinction of the non-avian dinosaurs
Fossils of dinosaurs stop appearing just above the Cretaceous-Paleogene (K-Pg) boundary, dated to just over 66 MYA. This is the most recent of the five mass extinction events in Earth’s history. Fossil record biodiversity shows a sharp drop in other species by 75%, with a simultaneous reduction in both marine and terrestrial environments.
The most widely supported explanation for the cause of the extinction is the impact hypothesis. The long line of evidence for this hypothesis includes:
- 200 km wide Chicxulub crater discovered by Alvarez and son in the 1970s, off the coast of the Yucatán Peninsula in Mexico.
- Granite found at the crater’s seafloor peak ring, which is usually only found deeper under oceanic crust, indicating uplift due to the impact
- Gravity and magnetic anomaly surveys confirmed the feature as an asteroid impact crater
- Impactite rocks (e.g. shocked quartz, suevite and tektite glasses) found at the site (DePalma et al., 2019)
- Fossil evidence of mass extinction of many species found in North Dakota at the K-Pg boundary, including fish filled with debris from the impact. Analysis of the fish bones shows the impact event occurred around springtime. (DePalma et al., 2019)
- Worldwide thin (~1 cm) layer of iridium-rich clay (160 times the usual concentration) found at the K-Pg boundary, indicating an extraterrestrial origin, since iridium is only found at high levels in asteroids (iridium is siderophilic; it naturally alloys with iron and sinks into the Earth’s core) (Goderis et al., 2021)
- Isotopic ratios of other transition metals found in the layer such as osmium, ruthenium and chromium closely match those found on carbonaceous chondrites (asteroids from the outer solar system) and not elsewhere in the crust (Fischer-Gödde et al., 2024)
- Two independent studies using argon-argon dating have obtained precise dates for the impact event of 66,043,000 ± 11,000 years ago and 66,051,000 ± 31,000 years ago, which are consistent with each other and the date of the K-Pg boundary itself.
Another hypothesis is the volcanic eruption of the Deccan Traps in India, which would have caused sudden climate change due to release of sulfur dioxide aerosols, suddenly dropping the temperature. However, most recent studies conclude that this was either merely a secondary factor, or that it was not a factor at all in the extinction.
Most of the world’s forests died off in the event, with pollen analysis finding that only two types of fern plants survived. The tree-dwelling birds all died out with the impact, and only a small number of land-dwelling birds seems to have survived as the ancestors of all modern birds (Waters, 2018). A small number of non-avian dinosaur fossils are found above the K-Pg boundary, implying that the extinction was a slow process due to the climate change following the asteroid impact, rather than the moment of the impact itself. The fungal infection mammalian selection (FIMS) hypothesis suggests that the subsequent rise of mammals was due to their greater resistance to fungal infections compared to reptiles (which require sunlight exposure) as cloud cover increased and pollen/fungal spores spread in the aftermath of the impact (Casadevall & Damman, 2020).
Radiometric dating and its verification
Radiometric dating
Uniformity of decay rate parameters
The laws of physics are observed to be uniform across space and time, and radioactive decay rates depend only on fundamental physics (gauge theory: nuclear forces and quantum field theory). The mechanisms of decay are sufficiently well understood (e.g. Gamow theory of alpha decay, and Fermi / Gamow-Teller theories of beta decay) that we can understand (and test) in exactly what conditions would be necessary to perturb decay rates.
Studies such as (Emery, 1972) investigated a wide variety of radioisotopes and stimuli (temperature, pressure, EM fields…) and showed that decay rates are immutable except for extremely minor changes and/or highly unnatural conditions due to well-understood physical mechanisms (e.g. electron capture cannot occur for fully ionised atoms since there are no electrons to capture). (Pommé et al., 2018) and (Kossert & Nähle, 2014) also found no dependence on decay rates by neutrino flux or solar output. Without any evidence for the catastrophic conditions necessary to perturb decay rates, we can be confident that decay rates have remained constant over geologic time, enabling reliable radiometric dating.
Radiometric Dating with Uranium series
With coral dating: here
Mount St Helens dating, using isochron dating
Argon dating of Mount Vesuvius
Many volcanic rocks naturally contain the isotope potassium-40 ($ ^{40}$K), which decays ~10% of the time to the stable isotope argon-40 ($ ^{40}$Ar) via electron capture followed by gamma decay with a half-life of 1.25 billion years. When these rocks first form from molten lava, any argon is expelled to the atmopshere on solidification, and the $ ^{40}$K begins to decay from its initial concentration at a predictable rate, forming trapped $ ^{40}$Ar in the rock.
Pliny the Younger was a Roman eyewitness to the Mount Vesuvius eruption, which he recorded as occuring in the early afternoon of 24th August, 76 AD, destroying Pompeii, Herculaneum and other Roman cities. In 1997, a piece of volcanic tephra from the region was subject to $ ^{40}$Ar/$ ^{39}$Ar (argon) dating, yielding an age of 1925 $ \pm $ 94 years: only 7 years older than the actual age of 1918 years. A second sample, sanidine phenocrysts in pumice, was taken in 2004 using the same method, which yielded an age of 1925 $ \pm $ 66 years, which is the exact calendar year. Additionally, in 2003, the same sample was dating using the U-Th/He dating method, giving an age of 1866 ± 243 years, which is very precise considering this method is usually used for dating much older rocks.
Carbon dating of the Teide volcano
The radioisotope carbon-14 is continuously formed in the upper atmosphere by cosmic rays, which can be absorbed by plant matter as CO$ _2$ (at a slightly lower $ ^{14}$C abundance due to slower gas diffusion of $ ^{14}$CO$ _2$) and incorporated into the plant’s tissues (e.g. glucose, cellulose). When the plant dies, carbon exchange stops, and the remaining carbon-14 decays with a half-life of 5,700 years. The ratio of $ ^{14}$C to $ ^{12}$C in a sample of organic matter can therefore be used to estimate the time of death of the organism, up to about 50,000 years ago due to resolution limits.
The Teide volcano in located in Tenerife (the Canary Islands). Stratigraphy found an age younger than 2000 years, while paleomagnetic dating found an age of 500 - 900 years. Historical records give an age of at least 500 years (European settlement began in 1494): Christopher Columbus reported seeing “a great fire in the Orotava Valley” as he sailed past Tenerife on his first voyage to the New World in 1492, interpreted to have been the Teide eruption. $ ^{14}$C dating gave a precise range of eruption between 1470 - 1490 AD (510 - 530 years ago). K/Ar dating gave a range of 800 $ \pm $ 300 years ago. These ranges and accounts all corroborate with each other.
Dating of recent fossils and artifacts
Cheddar man is a well-preserved skeleton of a Mesolithic (middle stone age) Homo sapiens found in the UK. DNA analysis found that he was likely a hunter-gatherer with bright blue-green eyes, slightly curly hair and black skin, with no lactase persistence. He probably arrived there via Doggerland, a low-lying region of Europe spanning between modern-day Britain, France and Germany, which sank under rising sea levels around 10-7 kYA, as the last glacial period ended. His Y-chromosomal haplogroup was I2a2, and 10% of British ancestry can be linked to Cheddar Man. Cheddar man was radiocarbon dated on two occasions to 8540-7990 BC and 8470-8230 BC, i.e. about 10,000 years ago.
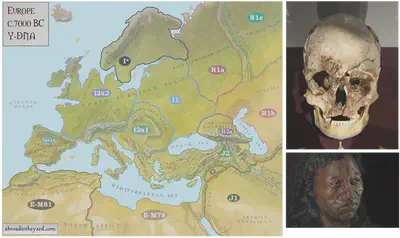
Ötzi the Iceman is a copper-age naturally frozen mummy radiocarbon dated to about 3,200 BC, found in the Alps. This is consistent with the materials, tools and food found with him, including a copper axe (at least 4,000 years old). DNA analysis found him to belong to Y-chromosomal haplogroup G2a-L91 (found today near the Mediteranean), and mitochondrial haplogroup K1f (extinct today). He had brown eyes, brown hair and a lactose intolerance. His genome is 99.7% identical to modern Europeans, with 5% of his DNA being Neanderthal.
Bog Bodies are a category of archaeological finds of naturally mummified human remains found in peat bogs, which preserve soft tissue and hair. A large number of these finds are known, with famous cases being the Tollund Man and the Elling Woman. The oldest known bog body so far is the Koelbjerg man, independently studied with carbon dating and pollen analysis to about 8,000 BC, or around 10,000 years ago. DNA and stable strontium isotopes extracted from the teeth reavealed his sex and birth region, belonging to the Maglemosian culture of Mesolithic northern Europe.
The Dead Sea Scrolls were found in 1947 in caves near the Dead Sea, and contained the earliest known records of the books of the Bible, written in Hebrew, Greek and Aramaic. Radiocarbon dating of the different scrolls gave dates from between 400 BC and 400 AD, which were mostly within 100 years of the dates estimated by analysis of the writing style (paleography).
The Shroud of Turin is a linen cloth with a distinctive imprinting that resembles the traditional face of Jesus Christ, which is said to have appeared on the cloth shortly after his crucifixion. However, spectroscopic analysis in 1978 suggested the imprint was painted on using a red ochre and vermilion pigment commonly used in medieval art. Additionally, in 1988, three independent radiocarbon dating laboratories all dated the cloth to between 1260-1390 AD, matching its first known appearance in church history in France.
The Vinland Map is a medieval map that allegedly shows the Viking discovery of North America, but was later found to be a forgery. Radiocarbon dating of the parchment and ink in 2009 found that the parchment was from the 15th century, while the ink contained modern carbon black, which was not used in medieval Europe.
Han van Meegeren was a Dutch painter during World War 2 and orchestrated a sophisticated art forgery. While his art skills were considered mediocre, he was able to create several convincing fakes of 17th century Vermeer paintings, some of which were sold to the Nazis in exchange for Nazi loot, including his piece The Supper at Emmaus. Van Meegeren confessed in 1946 and was found guilty with other evidence, but some doubt remained. In 1967, a study using the $ ^{210}$Pb radiometric dating technique was used to analyse the white lead (lead oxide) used in the paint. The smelting process to obtain lead removes much of the radium, which decays to $ ^{210}$Pb, so the method studied the ratio of $ ^{210}$Po (as a surrogate for $ ^{210}$Pb) to $ ^{226}$Ra. It was shown that the paint could not have been made more than a few decades prior (the 1930s), rather than the 300 years ago if it were genuine. Another study using gas chromatography in 1977 confirmed this finding.
The Voynich manuscript is a mysterious illustrated text with an unknown origin, language and purpose. Carbon dating of the vellum (animal skin) returned an origin between 1404 and 1438. It has been proposed to be written by 15th-century North Italian architect Antonio Averlino, consistent with the text and illustrations being all characteristically European.
Thermoluminescence dating
When radioactive decay occurs in crystalline minerals, the high-energy radiation can be absorbed by nearby valence band electrons, promoting them to the conduction band and leaving behind a hole in the valence band. Due to point defects in the crystal lattice, trap levels in the band structure capture these scattered electrons in a local metastable bound state, preventing recombination and fluorescence. Only when the crystal is heated (usually above ~500 C), these trapped electrons acquire sufficient energy to escape and recombine with the holes, releasing energy as photons called thermoluminescence (TL). The released TL photon intensity is proportional to the number of trapped electrons, which in turn is proportional to the amount of radiation (either by radioactivity or cosmic rays) absorbed by the crystal since its last heating event. By calculating the background radiation level per year from rates of decay and cosmic ray flux, the age of the crystal since its last heating event can be found.
In (Roberts, Jones & Smith, 1990), TL dating is corroborated with radiocarbon dating and used in tandem with artefact finds to date the first peopling of northern Australia to between 50 and 60 kYA.
TL dating is also used to date ceramics, tools and pottery. In (Rink & Bartoll, 2015), TL dating was used on the geometric Nasca stone lines in the Peruvian desert, finding them to have been constructed between 400 and 650 AD.
Electron spin resonance dating
Amino acid racemisation dating
Oxygen isotope ratio cycle ($δ^{18}O/δ^{16}O$)
The orbital monsoon hypothesis is based on the well-established concept of Milankovitch cycles, where long-term changes in Earth’s orbit (axial tilt, eccentricity and precession) result in changing frequencies, intensities and distributions of monsoons (intense wind and rain) on Earth due to changes in solar insolation. This has a strong impact on the climate of North Africa.
There are several lines of evidence for this model, including:
- The repeated occurence of sapropels (dark organic-rich marine sediment) in the plankton-rich sediment cores of the Mediterranean sea (low oxygen content due to freshwater influx from the River Nile).
- The repeated occurence of freshwater diatoms blown into the ocean sediment cores in the Atlanic ocean due to heavy monsoon trade winds.
- The cycle in isotopic ratios of oxygen-18 to oxygen-16 in calcite stalagtites/stalagmites in caves in China and Brazil, indicative of cycles in the water temperature due to the variable climate.
These cycles all have a period of about 22,000 years, closely matching the precession cycle of the Earth.
Sources: here (video).
Saltwater and freshwater have different ratios of oxygen isotopes, due to more evaporation (depletion of oxygen-16) in the ocean. This means that we can learn about what sort of water an animal drank by studying the isotopes that were incorporated into its bones and teeth as it grew: higher oxygen-18 content implies saltwater (marine), while lower oxygen-18 content implies freshwaster (rivers and estuaries).
The isotopes show that Ambulocetus (transitional whale) likely drank both saltwater and freshwater, which fits perfectly with the idea that these animals lived in estuaries or bays between freshwater and the open ocean. Whales that evolved afterwards (Kutchicetus, etc.) show even higher levels of saltwater oxygen isotopes, indicating that they lived in nearshore marine habitats and were able to drink saltwater as today’s whales can.
).](/post/evolution-evidence/whale_fossil_fresh_to_saltwater_hua5377fea055342a0c5ed01e61d45cbab_273508_8323fe1c252973ea6df9615bb0bf11fb.webp)
Ice core paleoclimate data
6. BIOGEOGRAPHY
Ecological succession
Primary succession describes the macroscopic sequence of events that follows formation of a new region of land (well-studied in physical geography) as life moves in for the first time. The resulting ecosystems that form (in the ‘climax community’) are highly interdependent, such that removing one would collapse the whole food web, which is a defining feature of irreducible complexity. Yet, we watch it happen all the time - the interdependencies are only ’locked in’ later on, not at the start.
Sources: here (article), here and here.
Prediction of Tiktaalik
Evolution had long predicted (since Darwin) that tetrapods evolved from lobed-finned fish, implying there should exist a range of fossils showing the transition from water to land across the Devonian period (about 400 MYA), with traits from both tetrapods and their fish ancestors. By the early 2000s, fossils including Eusthenopteron (385 MYA, a lobe-finned fish), Ichthyostega and Acanthostega (365 MYA, early tetrapods) had already been found, with the synapomorphies of labyrinthodont teeth and tetrapod skull roof pattern linking them together.
In 2004, Neil Shubin and his team predicted that there should exist a transitional fossil between these species, in late Devonian rocks (~375 MYA) in the Canadian Arctic. This location was chosen for its well-preserved sedimentary formations from freshwater and deltaic environments of the period, with the Fram Formation of Ellesmere Island being the optimal candidate. Using stratigraphy to identify the correct depth, Shubin’s team discovered three fossil specimens of the new species Tiktaalik roseae in 2004-2006, which collectively covered all of the species’ anatomical features.
Evolutionary pressures favouring land adaptation for these fish included:
- predation by the colossal Dunkleosteus in the deeper water, promoting survival in shallower waters near the shoreline
- regular desiccation in these shallower regions, requiring occasional survival out of the water
- ample untapped food supply of arthropods on land, rewarding those fish that could venture out of the water
The prediction and subsequent discovery of Tiktaalik is a powerful demonstration of the predictive power of evolutionary theory, and an example of hypothesis testing by observation, which are the hallmarks of a robust scientific theory.
7. COMPARATIVE ANATOMY
Body plans
Using the shared anatomy of each clade, as well as the fossil record, the characteristics of the basal ancestor of each clade can be inferred. These ancestors are visibly similar enough to also reasonably share their own common ancestor.
).](/post/evolution-evidence/mammal_body_plan_huc5c34a757af5b9ab144791f19121ba8d_1932244_99071ce05ae2ef60685660e7164c6e07.webp)
Middle ear bones in birds
.](/post/evolution-evidence/bird_ear_bones_1_hubc2eb35181f0972b2c749ada57ef817d_371485_278a5c02570e154f596b517b0a11d39d.webp)
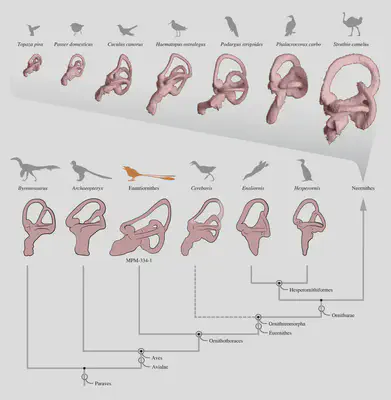
Nose position in whales
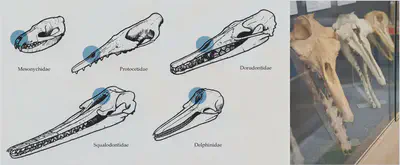
Limbs in fish and tetrapods
Homology increases with evolutionary relatedness. Using the well-known fish-to-tetrapod transitional fossil series, we can see how the bones in the fins of lobe-finned fish gradually evolve into the limbs of tetrapods, with the same arrangement of bones (humerus, radius and ulna) conserved throughout, but with increasing numbers of digits.
.](/post/evolution-evidence/bones_in_fish_huec3958c5c904a639800492aef2573234_1054979_71ad0a5ddbbf783dc9b46ab365d3ea21.webp)
Lungs and breathing in lobe-finned fish
As part of the water-to-land transition, the lobe-finned fish (Sarcopterygii) also had to adapt to breathing air rather than extracting oxygen from water via gills. Tiktaalik’s ribcage was imbricate (robust and with ribs overlapping), allowing expansion and contraction by muscles, implying they enclosed the lungs for breathing, unique among fish at the time. It also had otic notches, indicative of spiracles (gills like primitive blowholes, only found in air-breathing sarcopterygians), showing two modes of inspiration.
Limb attachment in lobe-finned fish
Eusthenopteron had pectoral (front-most) fins that connected to axial skeleton via part of the skull. In later species (e.g. Tiktaalik) and in tetrapods, this part of the skull is detached and comprises the pectoral girdle, also creating a neck for flexible head movement. A transition for this separation is seen in Panderichthys. It is known that bones form in development by two different mechanisms: 1) endochondral ossification (bone replaces cartilage) and 2) intramembranous ossification (bone forms directly from mesenchyme). The bones in the limbs of tetrapods are formed by endochondral ossification, while the bones in the skull and pectoral girdle (clavicle and most of the scapula) are formed by intramembranous ossification, demonstrating their former combined origin in lobe-finned fish.
](/post/evolution-evidence/limbs_in_fish_hu1828cea7216432f13d25f0f913c4cc45_566304_a7e8ad2585c476cf5ec9b86391dd7b5d.webp)
Limbs in vertebrates and mammals
When we zoom into the vertebrate clade, we see similarity increase further. All vertebrate animals share the same structures in their arms (humerus, radius, ulna, carpals, metacarpals, phalanges).
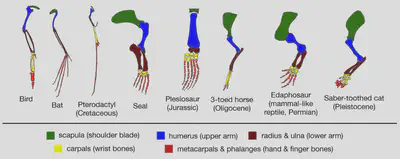
Within mammals, the similarities are further still, with all mammals sharing the same number and arrangement of the bones, but in variable lengths.
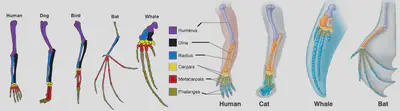
Reaction wood in trees
Trees must respond to deviations from vertical growth to avoid structural failure (thigmomorphogenesis). The group of woody plants, including trees, have evolved a specialised tissue called reaction wood, which is deposited at the base of the trunk in a way that counteracts the bending moment caused by gravity on a leaning tree trunk.
In all gymnosperms (softwoods, non-flowering plants), the reaction wood forms on the inner side of the leaning trunk (in compression), called ‘compression wood’ (high lignin content). Meanwhile, in angiosperms (hardwoods, flowering plants), the reaction wood forms on the outer side (in tension), called ’tension wood’ (high cellulose content). There is one exception to this trend: the clade Amborella, which is an angiosperm, but produces compression wood like gymnosperms. Amborella has been shown by molecular phylogenetics to be the most evolutionarily basal extant angiosperm lineage (i.e. it is the sister clade to all other flowering plants). This divergence pattern is therefore entirely consistent with the predictions of evolutionary theory: the parsimonious conclusion is that the tension wood trait evolved once in the lineage leading to all other angiosperms after their split from Amborella.
).](/post/evolution-evidence/tree_evolution_hue651627ee9189f6c4fda8110ae805083_138410_66bbb22662b63b047857475ba9cc2815.webp)
Reaction wood production is mediated by the plant hormone auxin (indole-3-acetic acid, which also controls gravitropism more generally) and ethylene, which act as signalling molecules to induce asymmetric cell growth in the cambium layer of the trunk. Mutations to the genes in these regulatory pathways can therefore alter the reaction wood response, potentially being responsible for the transition from compression wood to tension wood in angiosperms.
Living “transitions”
Some species that are alive today show interesting features that strongly indicate their evolutionary origin.
- Lungfish: have both gills and lungs, allowing them to survive in water and on land for short periods.
- Walking fish: fish such as the mudskipper and red-lipped batfish are poor swimmers (adapted for benthic regions) and use their adapted fins to ‘walk’ on the sea floor or on land for short periods.
- Duck-billed platypus: lays eggs like a reptile, but is a mammal (produces milk, has fur, is warm-blooded). Male platypuses have venomous spurs on their hind legs, similar to some reptiles.
The eye in vertebrates
Anatomical constraints in eye evolution: here
8. COMPARATIVE PHYSIOLOGY
Endosymbiosis in insects
Aphids have an endosymbiotic relationship with Buchnera aphidicola bacteria. The bacteria break down nutrients that aphids need to survive but can only live within specialised ‘bacteriocyte’ structures in Aphids. But some aphids later in their evolution dropped B. aphidicola and now have a yeast-like symbiont (YLS) that performs similar functions. These aphids still have the bacteriocytes but the YLS is located both inside and outside of them. So, these aphids have the same specialised structures as their cousins to host the bacteria but their symbiote is a fungus that doesn’t need those structures.
An example of higher-degree endosymbiosis is the Darwin termite (Mastotermes darwiniensis). The termite relies on a protist Mixotricha paradoxa to process the wood. The protists further rely on other bacteria living on its surface (each look like a thin hair that wiggles; symbiotic signalling in exchange for food). Within the protist, there is another endosymbiont spherical bacterium that digests cellulose in wood, releasing acetate for the protist. These bacteria have removed the need for M. paradoxa to have mitochondria, which have degraded into simpler organelles (hydrogenosomes and mitosomes).
Sources: here and Chapter 38 of The Ancestor’s Tale by Richard Dawkins.
An example of potential endosymbiosis occurring in real time is the obligate endosymbiotic gammaproteobacterium Carsonella ruddii, which lives inside psyllids (phloem sap-feeding insects). C. ruddii supplies the host with some essential amino acids, and has an extremely efficient genome consisting of 97% protein-coding DNA, with overlapping shortened genes in one of the smallest known genomes (182 genes in 159,662 base pairs, and getting smaller over time). Numerous genes considered essential for life seem to be missing, suggesting that the species may have achieved organelle-like status.
Evolution of eyesight
As one of the most impressively complex sensory organs, the eye’s evolution has been especially well studied. Eyes in some form have evolved independently over 50 times, with vastly different structures and functions suited to each lineage.
Different types of eyes
- Retinal phototrophy: the retinal molecule is used by Haloarchaea in bacteriorhodopsin proteins to capture light energy via chemiosmosis. This is the key light sensor molecule needed for eyes, without any of the complex structures that evolved later for information extraction rather than energy harvesting.
- Eyespots: a simple light-sensitive organelle found in euglenids (unicellular photosynthetic protists), containing photoreceptor proteins. Without any nerve cells, the signal cascade on light detection results in flagellar movement, enabling phototaxis.
- Developmental control genes: PAX6 is the master control gene for eye development in multicellular life, conserved across all animals (Gehring, 2002).
Animals have two main kinds of photoreceptor cell: ciliary (mostly in vertebrates) and rhabdomeric (mostly in invertebrates) and both kinds use retinal. Some animals have both, like the lab polychaete annelid Platynereis dumerilii.
- Pit eye: one of the types of eyes found in invertebrates. A pit with photosensitive cells inside allows for some vague directionality of light detection.
- Pinhole camera eye: Found in Nautilus. More directional sensitivity, by nearly closing the pit, allowing light to enter only through a small aperture.
- Lens formation: Evolved 8 times. An inhomogeneous lens formed of crystalline proteins continuously bends light for focussing of light onto the photosensitive layer, giving a clearer image. It also corrects for spherical aberration.
- Multiple lenses: Found in Pontella. Males have three lenses; females have two. The extra front-facing lens in males is parabolic and corrects for spherical aberration of the other 5 surfaces. The retina has only 6 receptors.
- Telescoping lens: Found in Copilia. Two lenses work like a telescope with a point-like retina and only a 3° field of view. There is a horizontal scanning eye movements at <5 Hz, while the bottom apparatus (eyepiece + retina) moves in image plane of the ‘objective’. The prey (plankton) moves vertically, giving a second dimension of scanning.
- Corneal refraction in land animals: to correct for the air-water interface and spherical aberration. In humans, 2/3 of the optical power is in the cornea rather than the lens.
- Reflective concave mirror (argentea) in the scallop.
- Tapetum lucidum: a reflective layer behind the retina in many nocturnal and crepuscular mammals (cats, dogs, deer), aiding in night vision. A variety of structures and molecules are used.
- Compound eyes in insects and crustaceans.
- Nanostructured cornea anti-reflection surfaces for quarter-wave matching in moths.
- Binocular/stereoscopic vision in vertebrates.
- Trichromatic vision in primates. The Old World monkeys (including apes), as well as some female New World monkeys, are trichromatic, having gained a third cone from the dichromatic mammalian ancestral lineage.
Physical constraints in the evolution of the eye
Solar radiation is the main source of high-exergy (low-entropy) free energy in the open biosphere, and its exploitation is therefore strongly selected for, such as in photosynthesis, photocatalysis and eyesight. At the molecular level, interaction with light requires a molecule that can absorb photons at the appropriate energy (wavelength), typically found in highly conjugated organic molecules. For wavelengths in the visible spectrum (most intense at Earth’s surface), these molecules include e.g. chlorophyll, retinal, 7-dehydrocholesterol, bacteriorhodopsin and phototropin.
With eyesight, there is the additional task of extracting information from the radiation, providing both thermodynamic and information-theoretic constraints on the evolution of the eye.
Radiation entropy maximisation: solar radiation is the main source of high-exergy free energy in the open biosphere, and its exploitation is therefore strongly selected for (e.g. photosynthesis). Solar radiation contains both energy and entropy, with slightly different Wien peaks for the two. The eye evolved to maximise the information extracted from the radiation, and the eye’s spectral sensitivity is optimised to the maximum entropy peak instead of the energy peak. Sources: (Delgado-Bonal & Martín-Torres, 2016) and (Delgado-Bonal, 2017).
Utility-based coding: updates the opponent process theory explaining how S, M, L cone signals are encoded in the optic nerve and visual pathway. The new theory describes the optimal encoding of spectral information given competing selective pressure to extract high-acuity spatial information. Source: (Conway, Malik-Moraleda & Gibson, 2023).
Principal components of reflectance spectra: the encoding of S, M and L into three channels can be explained by the observation that the three channels are the first three PCs in a PCA of the reflectance spectra of natural materials and scenes, encoding 98% of the total variance. This allows minimal information loss with the fewest number of channels, a consequence of the fact that the eye evolved to extract information from the environment. Source: (Chiao, Cronin & Osorio, 2000).
Evolution of photosynthesis
Photosynthesis is another form of phototrophy which uses chlorophyll as the light-absorbing molecule. Like the eye, it is extremely complex today, but started out from simpler systems based on the same fundamental principles.
Photosystems in unicellular life
The two main parts of photosynthesis are Photosystems I and II:
- PSI: an electron transport chain using ferredoxin to generate NADPH.
- PSII: a water-splitting complex generating protons, which can be used for chemiosmosis in ATP synthase. Likely to have evolved first due to its simpler structure and immediate utility in generating ATP.
The ATP produced can then be used in a metabolic cycle to fix carbon dioxide into useful organic compounds.
The bacterial kingdom Bacillati contains a range of photosynthetic bacteria:
- Phylum Cyanobacteria: contains both Photosystem I and II, using the Calvin cycle.
- Phylum Bacillota: only uses Photosystem I, without any associated synthetic cycle.
- Phylum Chloroflexota, order Chloroflexales, only uses Photosystem II, using the 3-hydroxypropionate bi-cycle.
The bacterial kingdom Pseudomonadati also contains a similar variety:
- Phylum Chlorobiota (green sulfur bacteria): contains Photosystem I, using the reverse Krebs cycle.
- Phylum Pseudomonadota (purple bacteria): contains Photosystem II, using the Calvin cycle.
Cyanobacteria became incorporated into eukaryotes as chloroplasts via endosymbiosis, allowing plants and algae to make use of photosynthesis, with both photosystems I and II.
9. DEVELOPMENTAL BIOLOGY
How is it that every cell in your body has the same DNA, yet different body parts can have completely different functions? How does your body know where to put everything? Development from an embryo is a tightly-regulated process, with the goal of controlling what genes get expressed where and when. There is a close relationship between evolutionary diversity and developmental diversity, and so we can study one to learn about the other. This is the basis of evolutionary developmental (evo-devo) biology.
Vestigial structures
Evolutionary change sometimes lags behind the needs of an organism. Body parts inherited from ancestors that had utility may become useless or serve a totally different function (exaptation) in a new environment. These are vestigial structures.
There are some structures that were considered vestigial in the past on the basis that they appeared to serve no function, but later scientific study found functionality. If there is no evidence for an evolutionary change in this function, then these structures are not vestigial (e.g. appendix, yolk sac…)
The vestigial tail and pharyngeal gill slits
Pharyngeal gill slits: all deuterostomes have gill slits in their pharynx, originally used for filter feeding, and evolving into gills in fish. They have been lost entirely in echinoderms. In mammals, they become inner ear structures and throat cartilage.
Post-anal tail: all chordates have a tail at some point in their life. In the more recent clade Hominoidea (apes), the tail has been lost, regressing normally by week 8 of development in humans. In very rare cases, mutations can upregulate the developmental gene Wnt-3a, whose normal suppression induces apoptosis in the cells of the tail, leading to the re-emergence of the tail at birth where it can be surgically removed (an atavistic trait).
). **(b)**: a rare disorder where the tail persists until birth (an atavistic trait) ([Shad & Biswas, 2012](https://casereports.bmj.com/content/2012/bcr.11.2011.5160)).](/post/evolution-evidence/human_embryo_tail_hu7e90ae0549284469bfc7623a9606abe3_657690_5d17b64a84a45410fcdf14be996d291c.webp)
Palmaris longus muscle tendon
The palmaris longus is a small muscle in the forearm that is absent in about 10-15% of the population, with negligible loss of overall grip strength or hand function (there is a small reduction in pinch strength in the 4th and 5th fingers). The rate of occurrence rises to over 50% in Egypt and Turkey (Ioannis et al., 2015). It is a vestigial tendon that controlled the retractable claws in ancient ancestral mammals.
Vestiges from recent hominin evolution
Other notable vestigial traits in humans include:
- Auricular muscles are Tiny muscles that move the ears and scalp, used for directional hearing in other mammals but too weak to be useful in humans.
- Goosebumps (arrector pili muscles), used to raise hairs for insulation and appear larger to predators in other mammals, but mostly ineffective in humans due to our relative hairlessness.
- The coccyx (tailbone), a remnant of the tail in our primate ancestors, which serves as an attachment point for muscles of the pelvic floor.
- The plantar reflex in infants, where the toes curl around an object placed under the foot. This is a remnant of the fully functional grasping reflex seen in primate infants, which helps them cling to their mothers.
Blind fish
The blind cave fish (Astyanax mexicanus) is a species for which some live in dark caves their whole lives, while others live above ground in rivers. The cave-dwelling fish differ significantly in their appearance: they have non-functional remnants of eyeballs (vestigial; sometimes fully lost), have completely lost the pigmentation in their scales (albinism, due to a mutated OCA2 gene) and they also have significantly smaller midbrains (the region of the brain that processes vision). These traits have convergently evolved independently in multiple cave populations.
Experiments have shown that the blind fish have a metabolic energy reduction of 15% relative to the above-ground fish (Moran, Softley & Warrant, 2015), providing a selection advantage for the cave fish to lose their eyes (the ’expensive tissue hypothesis’). The blind cave fish is likely on its way towards peripatric speciation.
Comparative embryology
Von Baer’s law of embryology refers to the observation that in the earlier stages of development, embryos of closely related animals are similar. Diversification of the embryo follows evolutionary diversification. This differs from Haeckel’s idea that “ontogeny recapitulates phylogeny”.
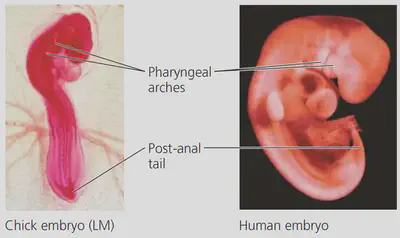
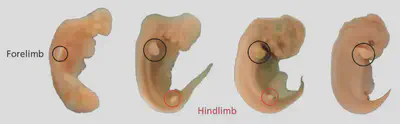
Evo-devo
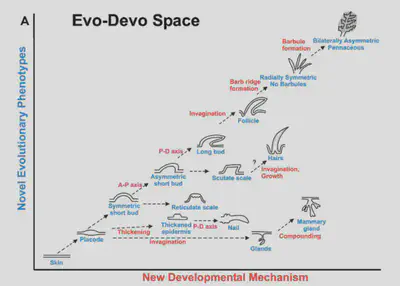
Homeotic genes: Hox, ParaHox, Pax, MADS-box
10. POPULATION GENETICS
Phylogenetic reconstruction
Comparative genomics studies the similarities and differences between the genomes of different species. This can be used to reconstruct phylogenetic trees, which show how closely related different species are. The more similar two genomes are, the more closely related the two species are likely to be.
A test of the validity of this reconstruction can be done using a known phylogeny. In 1992, a study was done on an artificially mutated strain of the virus bacteriophage T7, whose genome was sequenced repeatedly as it reproduced in bacteria. The experiment was stopped after 9 different viral strains had emerged, and only their genomes were used in 5 different phylogenetic reconstruction algorithms. All 5 algorithms produced the same, correct known tree, out of the 135,135 possible tree structures, with slight variation in the time to branching, showing that the algorithms are valid and can be used to reconstruct phylogenies from extant genome data more generally.
Source: (Hillis et al., 1991).
Great ape Y chromosome mutation rates
Sources: here
Statistical tests of common ancestry
(Baum et al., 2016) gives a rigorous and comprehensive statistical analysis of common ancestry applied to the primates. This paper is discussed here (video) and here (reddit post).
Maximum parsimony optimality criterion: a metric for scoring trees such that shorter trees (ones with fewer branching events) are considered more parsimonious, because these require fewer mutation events to explain the given genetic data. Tree length acts as a loss function to be minimised.
Characters and character states: a character is a categorical trait of an organism (e.g. hair colour), and the state of this character in a given organism is its value (e.g. blonde). In molecular phylogenetics, the characters are the loci of the DNA, and their states are the nucleotides identities.
Plesiomorphy and apomorphy: for a given clade, a plesiomorphy (ancestral trait) is inherited from an ancestor outside the clade. An apomorphy (derived trait) is evolved in some of the members of the clade.
Inferential statistics:
- A Monte Carlo method is a method wherein a computation of a deterministic quantity over an infeasibly large input state space is replaced with a simpler computation based on sampling from a random distribution over the search space, such that the Monte Carlo estimate converges to the true answer with more samples.
- A Markov chain is a random process over a defined state space for which, at each step of the process, the transition probability from one state to the next depends only on the current state (not on the past).
- A Markov chain Monte Carlo (MCMC) method is a Monte Carlo method where the target is the stationary distribution of a Markov chain. Such a Markov chain can be traversed, and the states visited are a representative sample of the distribution.
Permutation tail probability (PTP) test: a test of the explanatory power of a tree with taxonomic labels. Explained in (Archie, 1989) and (Faith & Cranston, 1991).
- For each character in the dataset, randomly shuffle the character states among the taxa.
- For the randomised data, compute the maximum parsimony phylogenetic tree.
- Evaluate the loss function (tree length).
- Repeat steps 1-3 several thousand times to generate a distribution of tree lengths $ X $ that are possible under randomisation.
- For the unrandomised data (i.e. the labelled tree being tested), find its most parsimonious tree length $ x $ (the test statistic).
- Calculate $ p = P(X \leq x) $ (the tail probability: the chance that random data could produce a maximum parsimony tree at least as short as the one being tested).
This an MCMC simulation of the distribution of tree lengths possible for a given randomised dataset.
For a tree containing genuine phylogenetic information, the PTP test will return an extremely small tail probability, since it should be improbable that such a tree could ever remain valid when the data is scrambled; random trees should not be able to compete with the real tree.
Baum et al.’s study: uses the genomes of a large sample of primate genomes (178 representative species) and identifies genes found to be conserved across the primates. The study uses this data to test three hypotheses individually:
- Common ancestry (CA) among all primates. Tested by running the PTP test on the labelled data as-is to generate a $p$-value.
- Family-level separate ancestry (FSA), where each family comprises a different tree. Randomly sample one member from each family and run the test on this data.
- Species-level separate ancestry (SSA), where each species is a whole ’tree’ (single branch)
For 1 (CA), the distribution was approximately normal and the test statistic was a minimum tree length of 10125. This is -71.8 standard deviations away from the mean of the distribution, which was 13422.3. This corresponds to $ p \approx 10^{-2581} $ (infinitesimally small). This is an overwhelming acceptance of common ancestry among primates.
For 2 (FSA, commonly proposed by creationists), the random distribution would have trees with correlated structures, at their starts, since they would be part of the distribution that had the most parsimonious trees. Assuming that a set of characters were not derived as a result of being in relation to another family does not mean that some family trees would not have a set of characters correlated or derived from some other means other than being related to another family tree. The actual data set of each actual known family fell way outside the generated distribution, so FSA is falsified.
The efficiency of MCMC in converging on an optimal solution by taking random steps through parameter space is powerful evidence for the strength and capability of the random mutation and natural selection mechanism to find the peaks of a fitness landscape: if the mechanism was ineffective, statistical parameter estimation methods wouldn’t work. This parallel is exploited in the success of evolutionary algorithms in computer science.
Peppered moths evolution prediction
Initial experiment in the 1800s, redone in the 1920s with a prediction, checked with genomes in the 1970s and verified.
11. METAGENOMICS
Great ape gut microbiome
Studies of gut bacteria in humans and other apes show that certain clades of microbes (Bacteroidaceae and Bifidobacteriaceae) have evolved along with their hosts for millions of generations. The timing of their genetic divergence matches the evolutionary split between humans and other apes, meaning that our gut bacteria, mitochondrial DNA, and nuclear DNA all diversified together. Some bacteria living in the human gut today are direct descendants of ancient symbionts that co-evolved and speciated in step with humans, chimpanzees, and gorillas, indicating our common ancient ancestry.
Source: here
Syphilis origin
The sexually transmitted infection (STI) syphilis (caused by the bacterium Treponema pallidum) originated in America about 9,000 years ago according to phylogenetic analysis. It only spread to Europe when Christopher Columbus arrived in 1492 and raped a large number of Native American women with his crew, who contracted the disease and brought it back. This is supported by the first historical record of syphilis in Europe being in 1495, when French troops invaded Naples.
Comparative genomic studies find that the syphilis-causing strain (T. pallidum pallidum) is a close sister to the yaws-causing strain (T. pallidum pertenue). A small number of mutations in genes controlling tissue invasion, immune evasion, and heat tolerance could have shifted it from a skin-to-skin childhood infection to an adult STI.
It has been proposed based on the archaeological record in pre-Columbian North America that the disease was less severe in its original form, occurring as a non-venereal treponemal disease (like those causing e.g. yaws). It quickly evolved into an extremely virulent and acute form in the high population-density and zero-immunity Renaissance-era European population, before ‘relaxing’ again to being a chronic STI as is known today (more balanced transmission and host survival).
Head, body and pubic lice in great apes
Humans have two types of lice: head/body lice (Pediculus humanus) and pubic lice (Pthirus pubis). Head/body lice are closely related to chimpanzee lice (Pediculus schaeffi), while pubic lice are closely related to gorilla lice (Pthirus gorillae). Phylogenetic analysis shows that the Pediculus lineage diverged from the chimpanzee lice about 6 million years ago, coinciding with the time of the human-chimpanzee split.
The Pthirus lineage diverged from the gorilla lice about 3.3 million years ago, indicating a host switch from gorillas to hominins (likely an australopithecine). It has been hypothesised that the host switch could only have happened after our ancestors had already lost most of their dense body hair, as otherwise the new lice would not have had an open ecological niche to occupy.
More recently, head/body lice Pediculus humanus later split into two ecotypes: head lice (living in scalp hair) and body lice (living in textiles of clothing). mtDNA analysis found that the body lice evolved <100,000 years ago, when humans began wearing clothes.
Sources: here (gorilla lice) and here (chimp lice).
Adaptation of the CRISPR-Cas9 system
https://enviromicro-journals.onlinelibrary.wiley.com/doi/10.1111/j.1462-2920.2007.01444.x
12. PHYSICS AND CHEMISTRY
Everything in biology is an emergent property of the underlying chemistry and physics, and so we can study the applications of these fields to biology to probe the fundamental principles evolution and life itself has to obey.
Bioenergetic penalty of non-functional genetic material
Every time a cell 1) replicates its DNA, 2) transcribes it into RNA and 3) translates it into a protein, there is an energetic cost to the cell, since these are anabolic (endergonic) reactions that require ATP consumption. An additional section of DNA (e.g. a new gene) therefore automatically consumes energy that would otherwise have gone towards maintaining the cell’s existing metabolism, comprising a potential loss of fitness for the organism.
In (Lynch & Marinov, 2015), it is shown that the cost of making a non-functional protein is typically above the selection threshold of natural populations, while the cost of retaining (copying) and transcribing the DNA into RNA (without translation to a protein) is typically below the selection threshold. This means that non-functional proteins are penalised by natural selection, but most non-coding DNA is not, and can accumulate neutrally by genetic drift. The new non-coding DNA can then potentially take on regulatory roles for the organism, or even undergo de novo gene birth into functional proteins.
The absence of an energy penalty to genome expansion is responsible for the ‘C-value paradox’ (genome size distribution is completely uncorrelated with evolutionary relatedness) and is exhibited by many plants which show polyploidy.
Thermodynamic permissibility of life and evolution
In thermodynamics, a ‘system’ is a specified volume of space with a boundary, whose energy and matter content can be quantified. The three types of systems are:
- Isolated system: ❌ energy exchange; ❌ matter exchange.
- Closed system: ✅ energy exchange; ❌ matter exchange.
- Open system: ✅ energy exchange; ✅ matter exchange.
For example, the Earth is (roughly) a closed system, since it receives energy from the Sun (insolation) and radiates thermal energy back out to space, but the mass transfers (atmospheric gas escape, space dust infall, mass defect from radioactivity) are negligible (sources: here and here).
The biosphere, and life itself (such as a cell) is an open system, and one in a highly non-equilibrium state, using free energy inputs to maximally generate entropy in the surroundings while maintaining a low-entropy internal state. The concept of exergy is useful in quantifying the potential of a closed or open system to do useful work and reduce its internal entropy. The exergy content of an energy source is the maximum useful work that can be extracted reversibly from it while rejecting heat to the surroundings, and the exergy of a system is given by
$$ B = H - T_0 S $$
where $ H $ is the enthalpy, $ S $ is the entropy and $ T_0 $ is the ‘dead state temperature’, which is the temperature of the surroundings where heat can potentially be dissipated to. Exergy can be used to combine the first law (conservation of energy) and the second law (entropy production) into a single equation, called an exergy balance.
For example, although a plant radiates away as much energy as it receives (otherwise it would heat up), the exergy of the incoming solar radiation is much higher than that of the outgoing thermal radiation, due to the Sun’s high blackbody spectrum temperature giving it a low entropy. The net exergy flux into the plant is what allows its internal processes to do useful work, which is to perform the endergonic, entropy-reducing reactions in photosynthesis. The transpiration stream of the plant provides the main high-entropy output of a plant in the form of water vapour, increasing the entropy of the surroundings significantly, allowing plants to harvest the external free energy of sunlight to create internal order. All life indirectly enjoys this benefit, since plants act as producers, providing energy (released by metabolism of food intake) for organisms at higher trophic levels in the food webs.
In Monod’s (1965 Nobel Prize in Physiology) 1971 book, Chance and Necessity, he recounts an experimental verification that life does not violate the 2nd law of thermodynamics:
We take a milliliter of water having in it a few milligrams of a simple sugar, such as glucose, as well as some mineral salts containing the essential elements that enter into the chemical constituents of living organisms (nitrogen, phosphorus, sulfur, etc.). In this medium we grow a bacterium, for example Escherichia coli (length, 2 microns; weight, approximately $ 5 \times 10^{-13} $ grams). Inside thirty-six hours the solution will contain several billion bacteria. We shall find that about 40% of the sugar has been converted into cellular constituents, while the remainder has been oxidized into carbon dioxide and water. By carrying out the entire experiment in a calorimeter, one can draw up the thermodynamic balance sheet for the operation and determine that, as in the case of crystallization, the entropy of the system as a whole (bacteria plus medium) has increased a little more than the minimum prescribed by the second law. Thus, while the extremely complex system represented by the bacterial cell has not only been conserved but has multiplied several billion times, the thermodynamic debt corresponding to the operation has been duly settled.
Useful resources:
- An intro-level explainer on thermodynamics in biochemistry, here
- Entropy and evolution (Styer, 2008), a basic pedagogical paper explaining why evolution does not violate the 2nd law.
- A YouTube video by Veritasium on the concept of entropy, with a section on entropy in life from 17:08.
- A practical thermodynamic analysis of photosynthesis (Keller, 2013).
- A rigorous exergy analysis of photosynthesis (Petela, 2008), with an emphasis on quantifying the exergy of sunlight.
- Biological catalysis of the hydrological cycle: life’s thermodynamic function (Michaelian, 2012), a comprehensive paper analysing the processes of life and solar radiation on the entropy and energy budgets of the Earth.
Non-equilibrium thermodynamics: the driving force for life
While elementary thermodynamics tells us that the processes of life are possible, it does not tell us how likely it is to occur, or how it actually happens. The more modern field of non-equilibrium thermodynamics is more suited to understanding the physical processes underpinning life, since all biological systems are open systems far from equilibrium. For any system subjected to a strong gradient of free energy, non-equilibrium thermodynamics tells us that the system will spontaneously self-organise into a state of maximum entropy production, known as a dissipative structure. For example, when a closed system of liquid water is heated strongly with a good-conducting boundary, it naturally forms turbulent convection cells which increases the mixing of hot and cool regions of the fluid, increasing the heat transfer rate and the irreversible entropy production rate in the system.
Useful resources:
- A YouTube video by NanoRooms on dissipative structuring in life.
- Life as a manifestation of the second law of thermodynamics (Schneider & Kay, 1994), a landmark paper establishing that life arises by necessity of thermodynamic principles, not just by possibility.
- Maximum power in evolution, ecology and economics (Hall & McWhirter, 2023), a discussion of Lotka’s statement of the ‘4th law of thermodynamics’ as it applies to evolutionary biology.
Some of the key authors in non-equilibrium thermodynamics include:
- Erwin Schrödinger, who wrote the book What is Life? in 1944, kickstarting the discussion of thermodynamics in the context of biology.
- Ilya Prigogine, who won the 1977 Nobel Prize in Chemistry for his work on dissipative structures and self-organisation, and wrote the textbook on the topic.
- Jeremy England, who applies dissipative structuring to the origin of life.
- Karo Michaelian, who has written extensively on the thermodynamics of evolution and abiogenesis. Some of his papers are featured in my ‘Origin of Life’ bibliography.
Information theory and evolution
“There are difficulties in applying information theory in genetics. They arise principally, not in the transmission of information, but in its meaning” (Maynard Smith, 2000, p. 181).
In the view of (Jaynes, 1957), thermodynamic entropy $ S $, as explained by statistical mechanics, should be seen as an application of Shannon’s information theory: the thermodynamic entropy is interpreted as being proportional to the amount of further Shannon information needed to define the detailed microscopic state of the system, that remains uncommunicated by a description solely in terms of the macroscopic variables of classical thermodynamics. This is captured in Boltzmann’s original formula $ S = k_B \ln \Omega $: the information entropy of a system is the amount of “missing” information needed to determine a microstate, given the macrostate. However, statistical entropy and thermodynamic entropy are not equivalent when applied to the genetic code: statistical entropy quantifies the uncertainty associated with an observation from a given distribution of possible sequences, while thermodynamic entropy is a property of a given individual sequence (and the physical interactions it has in vivo).
The data processing inequality (DPI) can be considered the information analog of the 2nd law of thermodynamics: if $ X \rightarrow Y \rightarrow Z $ forms a Markov chain, then no processing of $ Y $ (deterministic or random), can increase the information that $ Y $ contains about $ X $, i.e.
$$ I(X; Y) \geq I(X; Z), \ \ \ \ \ \text{where} \ \ \ I(X; Y) = H(X) - H(X | Y) $$
($ I $: mutual information, $ H $: information entropy.)
The DPI applies in the information analog of a closed system, where all of the influence of $ X $ on $ Z $ flows through $ Y $, and that there is no external intervention or feedback. Intuitively, evolution involves an increase in the raw information content of a genome, enabled by the fact that evolution is an informationally open system with feedback from the environment (natural selection). The mutual information with the environment must increase more than the entropy of the genome decreases:
$$ -\Delta H(\text{genome}) \leq \Delta I(\text{genome}; \ \text{environment}). $$
Random mutations tend to increase genomic entropy, while natural selection reduces it, allowing an increase in Shannon information (reduction in sequence diversity, increasing the data compressibility) if natural selection is strongly purifying (mutational effects are above the selection threshold). Natural selection can be thought of as a process that sorts through the genetic noise produced by random mutations, looking for the signal of genetic variations that increase differential reproductive success.
Some examples of quantifying the Shannon information gains occurring in evolution are given below:
In Chapter 19 of Information Theory, Inference and Learning Algorithms by David MacKay, a model for the Shannon information of a simple binary coded genome is presented, with the conclusion being that meiosis enables the information content of a genome to grow much faster (due to recombination) than mitosis alone.
In (Vopson & Lepadatu, 2022), the informational (Shannon) entropy $ H $ of the genome of the SARS-CoV-2 virus (COVID-19) over its mutational history between 2020 and 2022 is calculated, and found that the processes of mutation and natural selection caused the Shannon entropy of its genome to continually decrease. This represents an increase in the Shannon information in the genome, as purifying selections acts on the virus. Although the paper oversteps by proclaiming a universal ‘second law of infodynamics’ (the observed entropy reduction may not have occurred had the mutational effects been below the selection threshold), the data does clearly show the effects of natural evolution on Shannon information. I left a technical comment on PubPeer criticising the paper’s overreach here.
13. APPLICATIONS OF EVOLUTION
This section focusses on practical applications of the evolutionary theory, primarily in engineering, medicine and agriculture. It does not include applications of explaining aspects of biology itself, which are numerous. The utility of evolution serves as a ‘proof of concept’ that the theory aligns with reality.
Protein Engineering
We can develop entirely new enzymes using ‘directed evolution’ of proteins with a variety of uses. By artificially cloning the gene for an enzyme and introducing mutations, screening for activity and stability, we ‘artificially select’ more optimal enzymes for our use case. This is a well-established lab procedure that puts evolution into practice.
Glucose biosensors: when type 1 diabetics need to monitor their blood glucose levels to time insulin injections, they use a glucose biosensor, which typically works by measuring the rate of reaction of a glucose-binding enzyme. The wild-type enzymes found naturally are usually not stable enough for reliable operation in a biosensor (due to activity reduction on immobilisation, stability to temperature variations and cofactor dependence), so new enzymes are needed, which come from directed evolution. These are the most common type of commericalised glucose biosensors in use today. Although enzyme-free biosensors have been researched, they have not yet been commercialised due to lacking the chemoselectivity that enzymes excel at, so evolution-backed biosensors remain the state-of-the-art in type 1 diabetes management as of 2025. (Gutierez et al., 2013).
Lactate biosensors: another type of electrochemical biosensor making use of directed evolution. Lactate biosensors are used by high-performance athletes to monitor lactate levels in the blood and sweat, indicative of fatigue, and require directed evolution for thermostability. They are also used in hospitals at triage to test for septic shock or risk of meningitis. (Minagawa, Nakayama & Nakamoto, 1995) and (Minagawa et al., 1998).
Artificial metalloenzymes: new enzymes can be designed to catalyse completely new chemical reactions unseen in biology. In one study (figure below), a new enzyme was evolved containing an abiotic di-rhodium cofactor, which catalyses the cyclopropanation of styrene and diazo compounds with enantioselectivity. Industrial applications include the synthesis of pharmaceutical (e.g. sulfonamide antibiotics), improvement of enzymatic biofuel cells, carbonic anhydrase-based carbon capture technology and bioremediation by degradation of toxins (PAHs, PCBs, organophosphates…). (Reetz, 2011).
 in the enzyme. Source: [here](https://www.nature.com/articles/nchem.2927).](/post/evolution-evidence/directed_evolution_hucca222e264223908c01fb941179ae07c_624275_e9387723610ac512bec29ffc5a10636b.webp)
Bio-detergents: subtilisin is a protease used in laundry detergents, which was engineered to be more stable at high temperatures and alkaline pH. Its cofactor dependence on calcium ions was also removed, as calcium would be chelated by EDTA in the wash. (Yang et al., 2018).
High-fructose corn syrup: glucose isomerase is an enzyme used in the production of high-fructose corn syrup (HFCS), which is sweeter than regular corn syrup. It was engineered to be more stable at high temperatures and to have a higher affinity for glucose. This allows for more efficient conversion of glucose to fructose, resulting in a sweeter product. Source: here.
Genetically engineered bacteria: E. coli can be engineered to produce new useful products, such as carotenoids. They can also produce bioplastics, which are biodegradable, although issues of scale-up and durability remain. (Schmidt-Dannert, Umeno & Arnold, 2000).
Bioremediation: engineering of bacteria extends to waste degradation in an effort to combat environmental pollution.
Bacteria have been observed to have evolved enzymes to degrade man-made PET (polyethylene terephthalate) plastics (Alam et al., 2025, news article here), both by selection in the environment of a Japanese recycling plant and in the oceans due to plastic pollution. The PET hydrolase enzymes identified carried the ‘M5 motif’ in their tertiary structure, which is a modification of the ‘DLH domain’ (a generic α/β-hydrolase fold found in many hydrolase enzymes) including a stabilising internal disulfide bridge. PETases have been explored for bioremediation purposes (source: here).
PFAS (per- and poly-fluorinated alkyl substances) are a now-infamous group of environmental pollutants commonly known as ‘forever chemicals’: organic molecules whose structures render them practically inert to biological and chemical attack. The carbon-fluorine bond is rare in biochemistry, with only a few dehalogenase enzymes known in organohalide-respiring bacteria. (Yang & Liu, 2025) reviews the very recent literature exploring the new possibility for bioremediation of PFAS in the environment. In (Jaffé et al., 2024) and (Huang et al., 2024), it is found that Acidimicrobium sp. strain A6 is capable of degrading PFOA, PFOS, and PFHxS using its rdhA gene. Another study, (Wijayahena et al., 2025), finds similar results for PFOS and some others by Labrys portucalensis F11. Unlike nylon, it is improbable that nature alone would evolve an enzyme to metabolise PFAS due to the recency, chemical dissimilarity and lack of environmental pressure, so technological innovations are needed. It is hoped that bioengineering techniques such as genetic engineering and directed evolution could help improve the rate and efficiency of PFAS degradation. While other solutions involving PFAS-absorbing probiotics (Lindell et al., 2025) have been proposed, environmental cleanup remains a challenge.
Animal Model Selection in Translational Medicine
The animals used in lab studies for medicines are chosen based on evolutionary relatedness. They use rats for most in vivo studies since they’re one of the closest non-primate animal orders to us (order Rodentia). Rabbits are in another very close order (order Lagomorpha), and we are all mammals.
For neurological studies, primates are sometimes used, as their brain structure is closer to ours. Improved animal welfare laws mean that primate studies are now rare (they are only done for behavioural studies and occasionally for neurosurgery, e.g. neural prosthetic implants), and some countries (e.g. the UK) are phasing out animal studies entirely, replacing them with modern biotechnologies such as organ-on-a-chip and 3D bioprinted tissues. However, as of 2025, animal studies remain highly important to modern medicine. Without evolution, we’d be stabbing in the dark as to whether a particular animal would serve as a good model for our in vivo testing, which would mean significantly fewer successful drugs and therapies passing the animal testing phase.
Importantly, medicine is complicated, and evolution is not the only factor in determining the translatability of animal model results to humans. To decide whether “X happens in animals ⇒ X happens in humans”, we must check:
- Is the causal mechanism evolutionarily conserved?
- Is the regulatory network conserved, not just the components?
- Are physiological differences (size, metabolism, lifespan) irrelevant to the mechanism?
- Does the animal model have construct validity (shared mechanism), not just surface similarity?
Only if all four conditions are met can you reasonably extrapolate animal model results to humans. In some diseases where there are relevant genotypic differences, mouse models are not effective representatives of humans, most notably cancer. Most cancer treatments animal-tested in mice fail in humans (more so than other drugs). In mice, telomeres are longer than in humans due to stronger telomerase activity, which reduces the contribution of cellular senescence to age-related cancers, making them less relevant for human studies. Telomerase repression is an evolutionary control measure for high body-mass mammals that compensates between longevity and cancer risk (Gorbunova & Seluanov, 2008).
Another example is the fact that heart xenotransplants predominantly use pigs as donors rather than chimpanzees, despite the latter’s closer relatedness, for three major reasons:
- Chimps carry viruses that are deadly to humans, e.g. SIV.
- Chimp hearts are also too small for human patients, as we are endurance-adapted with large cardiac output while chimps are not.
- Chimps are prohibitively impractical and unethical to keep in captivity for organ donation.
Chimps in general are rarely useful models for humans despite their high genetic similarity, due to a variety of epigenetic differences (Bailey, 2011). Pigs happen to have the right sized hearts and are far easier to domesticate. Genetic engineering of the pigs is required to remove the proteins that would trigger an immune rejection in humans: all xenotransplants without such genetic modification have failed (Wang et al., 2022).
Protein Folding Prediction
Protein structure prediction is famously hard task, and has only recently become feasible with powerful machine learning models like AlphaFold, trained on structures painstakingly obtained manually via cryo-electron microscopy and X-ray crystallography. AlphaFold uses a transformer-based ML architecture (the same structure as used in LLMs like ChatGPT) called the EvoFormer, which combines protein sequence data with data on sequence identity conservation across evolutionary lineages, which essentially provides information on which amino acid residues are crucial to the 3D structure and which are less constrained.
It’s hard to understate how revolutionary solving protein folding has been: it’s already been used to develop lots of new medicines by predicting protein-substrate interactions, and the newest model AlphaFold 3 can handle protein-DNA interactions too. AlphaFold 3 has recently been used to predict the consequences of how a virus will mutate during a pandemic which could help develop more robust vaccines. That’s using evolution to fight evolution! (Wee & Wei, 2025).
Universal Flu Vaccine
Influenza has been around since antiquity, but the most famous outbreak was the Spanish Flu pandemic (1918-1919, 1/3 world infected, 50 million deaths). Influenza A has 8 segments of ssRNA in their capsid, with two glycoproteins (envelope spike proteins): neuraminidase (N) and hemagglutinin (H). There are several isotypes of the H and N proteins, denoted H1, H2…, N1, N2… . Only H1-3 and N1-2 are human transmissible (at present!). The swine flu epidemic of 2009 was due to reassortment (genetic mixing) of the H1N1 bird flu, the H1N1 swine flu and the H3N2 seasonal human flu. The virus’ fitness is a balance of H and N expression, as too much of either will limit the lifetime of the virus. Anti-influenza drugs (e.g. Tamiflu) inhibit N activity, but must be taken early after infection.
Due to antigenic drift (spike protein point mutations allowing evasion of antibodies), flu is a seasonal virus. The DNA polymerase within influenza A is error-prone (lacks proofreading) and therefore mutates rapidly, maintaining its virulence by the high viral loads. Influenza usually has a low mortality rate, and most of its deaths are due to secondary bacterial infections (e.g. pneumonia) which occur when the lung tissue is compromised. If two different influenza virus subtypes infect the same cell at the same time, their individual ssRNA segments may form a ‘hybrid’ virus during capsid assembly (antigenic shift), which is how influenza can jump between species.
The PB1 protein in influenza A steals the protective 5’ caps from host nuclear mRNAs and uses them on its own RNA, both increasing its virulence and dampening the host’s self-protein synthesis and immune response. An alternate reading frame intrinsically disordered protein, PB1-F2, also contributes to virulence by inducing apoptosis in infected cells. A single point mutation, Asn66Ser, in PB1-F2 is associated with signfiicantly stronger pathogenicity and a more aggressive influenza infection.
A bird flu (H5N1) epidemic occurred in 1997. Since 2020, there has been an ongoing pandemic of H5N1, due to reassortments from a variety of flu strains increasing virulence among domestic poultry, and the virus is primarily responsible for the US chicken egg shortage in 2025. There have been a small number of human cases, though human-human transmission has not been observed as of 2025. The neuraminidase spike protein has been mutating over time, with longer alpha-helix stalk structure associated with increased transmissibility across mammalian species, and the rapid mutation rate of the H5N1 spike proteins make vaccine development challenging. A potential universal vaccine approach instead targets the nucleoproteins inside the capsid e.g. PB1, which are less tolerant to mutations, and can be recognised in infected cell MHC class I molecules by T cells. Universal bird flu vaccines are currently under development in case of a future human-transmissible bird flu pandemic.
- How evolution explains virulence: here (video)
- Explanation of bird flu and the universal vaccine: here (video)
- Development programme for universal flu vaccines (April 2025): here
Cancer Research
The evolutionary concept of virulence extends to cancer cells, with additional complications due to multi-level selection: cancer breaks the altruistic nature of multicellularity, placing them in direct competition with the host. Cancer’s trajectory is only explainable via evolutionary dynamics. It behaves like a pathogen where intra-host competition is maximised and inter-host competition is absent.
In her 2020 book Rebel Cell: Cancer, Evolution and the Science of Life, biomedical scientist Dr Kat Arney (PhD from Cambridge, working for Cancer Research UK) writes about how not recognising the evolutionary processes involved in cancer has held the field of cancer therapeutics back for decades, and we’re only now just catching up to break the stagnation.
Chemotherapy or radiotherapy are ‘brute force’ approaches that can kill as many healthy cells as cancer cells. Additionally, the ‘gene targeted’ approaches developed in the 2000s are ineffective as the cancer cells often mutate to resist the drug, and the cancer returns at a later date, so that further treatments of the same drug simply select for the resistant cancer cells. An adaptive therapy strategy is needed, modulating doses to force the cancer cells to compete with each other, rather than growing unimpeded. The tumor microenvironment behaves as an ecological niche which can be manipulated. This being incorporated into modern cancer therapies. Some of the treatments that can be/have been augmented with evolutionary principles to yield improved prognoses include:
- Mitotic checkpoint gene transfection, via mRNA vaccines: intended to control the cell cycle of cancer cells.
- Oncolytic virus therapy: immunotherapy, triggering an immune response. Effective in immune-rich tissues.
- Dendritic cell vaccines: another immunotherapy approach. Personalised medicine is required to moderate the immune response to avoid autoimmunity.
- Combination chemodrug-loaded hyaluronan hydrogels: cancer cells are less likely to evolve mutations to multiple bio-orthogonal drugs when delivered at once, reducing the risk of resistance. Using hyaluronan to target CD44 receptors increases specificity to leukemia stem cells. However, toxicity is also additive in combination drugs.
More info:
- How evolution explains cancer: here (video #3 in playlist)
- Dr Kat Arney discusses her book Rebel Cell: here
- Approved evolution-based treatment regime for prostate cancer: here
- Hyaluronan for CD44 targeting in leukemia: here
Basin Modelling for Oil and Gas Exploration
Basin modelling is a technique widespread in the global multi-trillion-dollar oil and gas industry, which synthesises geological, petrological and paleontological data to predict the locations of oil and gas reserves within the Earth’s crust. Common targets include oil shales from the Cambrian, Ordovician, Devonian, and Jurassic (as source rocks), as well as tight oil reservoirs found in Devonian, Carboniferous, and Cretaceous formations. It makes extensive use of radiometric dating and stratigraphy (e.g. foraminifera (protist) biostratigraphy) to date the sedimentary layers and model the thermal history of the hydrocarbon-bearing rocks.
In oil and gas, predictions mean profits, and errors mean tremendous financial losses! The success of this industry (at the expense of the climate, unfortunately…) would not be possible without the validity of the underlying theory.
There exists only one oil prospecting company in the world that refuses to use old-earth evolutionary models in their work: they are Zion Oil and Gas Corporation (ZNOG), founded by Christian fundamentalists who believe that Israel would yield oil reserves on theological grounds. Zion Oil has failed to find any ’economically recoverable’ oil reserves in over 20 years of trying, operates incurring annual losses of several tens of millions of USD and are practically bankrupt as of 2025, staying afloat only by selling shares to gullible investors. Sources: here and here.
Mining Exploration
Radiometric dating has also been used by mining companies. For example, in 2021, Hannan Metals used U-Pb dating of zircons to expand a Miocene-epoch deposit of gold and copper in Peru. Source: here.
Historically, American coal miners in the 19th century would use fossils of Archimedes (a Paleozoic-era bryozoan with a distinctive screw-shaped stalk) to indicate how to deep to dig when searching for coal. These bryozoans are commonly found in limestones that were older (lower in strata) than the mostly siliciclastic coal-bearing rocks and do not contain much of any coal, so would be pointless to excavate. (Jackson, 2008).
Evolutionary Algorithms
Evolutionary (genetic) algorithms are rarely the best solution to a problem since they are stochastic and highly generalised: they are essentially biased parallelised random searches, and this goes some way to explaining why biological evolution is so slow and messy. However, with domain-specific knowledge and a well-designed objective function and genetic encoding scheme, they can be very powerful, outcompeting conventional algorithms in some cases, illustrating the power of the underlying Darwinian principles.
The basic concept was explained by Dawkins in The Blind Watchmaker (1986), where he demonstrated how a random string of letters could be ’evolved’ into a target phrase (e.g. METHINKS IT IS LIKE A WEASEL) by iteratively mutating and selecting the closest matches. While the number of possible letter combinations is extremely large ($ 27^{28} \approx 10^{40} $), the evolutionary algorithm rapidly converges in on the order of 100 generations.
.](/post/evolution-evidence/darwin_evolutionary_algo.gif)
Evolutionary algorithms applied to various engineering problems include:
- Topology optimisation - Evolutionary Structural Optimisation (ESO) and ‘Generative Design’ in Autodesk Fusion 360. Examples in the review (Belhocine, Shinde & Patil, 2021).
- Hyperparameter optimisation in neural networks (Pujol & Poli, 1998) and (Jaderberg, 2017) from DeepMind.
- Training neural networks for deep reinforcement learning (here), from Uber’s research team, also (Zoph & Le, 2016) from Google Brain, and this post from OpenAI.
- Diffusion models are a type of generative AI used to create images or videos, and have been mathematically shown to be equivalent to an evolutionary algorithm (Zhang et al., 2024).
- Operations research (e.g. facility layout design, supply network design, vehicle routing, capacity planning, inventory management, scheduling, risk management) (Lee, 2018).
- Inverse problems, such as image reconstruction from PET or EIT data, which are used in medicine, soft robotics and particle technology (Abouhawwash & Alessio, 2021).
- Evolved antennas - Space Technology 5, and using COMSOL here
- FPGA circuit design (Ashraf et al., 2012).
Artificial Selection, Domestication and Genetic Engineering
Artificial selection is essentially to natural selection but with humans deciding what is ‘most fit’, widely used in agriculture and animal husbandry. We can guide the evolutionary process towards exhibiting desired traits using selective breeding.
Artificial selection has a tendency to evolve animals into phenotypic extremes that would not be good strategies in nature, due to the difference in selection objectives. This is the case for many dog breeds, horses and cattle. Artificial selection is a type of ’truncation selection’ since it acts on ‘finished’ phenotypes (e.g. milk output), and is therefore not equivalent to natural selection, which acts on all life stages from zygote selection to post-reproduction (in the case of helping the young).
Fruits and Vegetables
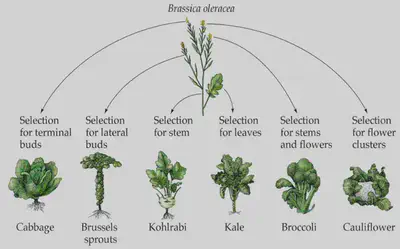

Animals
- Cats
- Dogs
- Livestock (cows, sheep, pigs, chickens)
- Pigeons: originally domesticated from the rock dove (Columba livia), but became suited to urban city environments as building roofs mimick their natural cliff-edge nesting habitats.
- Horses: pre-domestication horses were slightly smaller, and these wild horses are endangered today.
- Raccoons: urban raccoons have smaller snouts compared to their rural raccoons, a similar trait observed in domesticated foxes. Source: here (article) and here (paper).
Genetic Engineering and Gene Editing
- Gene drive for mosquitoes to eliminate malaria
Universal Darwinism
While “evolution” generally simply means “change over time”, the more specific meaning in biology can be generalised. Any imperfectly self-replicating system (subject to random variation with some mechanism for selection of survival based on that variation) can be said to evolve: this is the concept of Universal Darwinism, and it can be used as a philosophical or metaphysical framework to understand the dynamics of various other phenomena.
Chemical evolution refers to self-replicating molecules (autocatalytic systems) that can change their structures over time, which is especially relevant to the origin of life. Lone biomolecules like RNA are not alive and are therefore not part of biological evolution, but can still undergo this more general notion of ‘chemical evolution’.
Richard Dawkins extends evolution to the development of cultures, a concept called memetics, where ideas and cultural practices are the ‘genes’ that are subject to selection and variation, and can evolve over time.
X. EVOLUTION DEBATE
This section shifts from defending evolution to attacking its critics. These are not necessarily evidence for evolution, but are tangentially related and generally support the theory: useful to know as citations and rhetorical points in debates.
Scientific Consensus
The overwhelming majority of scientists accept evolution as the best explanation for the diversity of life on Earth. It is not an ‘argument from authority’ fallacy to cite the scientific community for a scientific argument, as the consensus is based on the evidence acquired through the scientific method, not on opinions or beliefs of individuals with power.
According to Pew Research Center, as of 2019, 98% of scientists accept evolution, whether religious or not. This percentage is higher than:
The status of a scientific theory is completely independent of the general public’s level of support for it. For example, among the American public (sources: here, here and here)
- 94% believe smoking causes cancer
- 84% (of Americans) believe the Earth is round
- 83% believe vaccines are safe and effective (in 2014)
- 60% believe the Earth is about 4.5 billion years old
- 58% believe in evolution (whether naturalistic or theistic)
- 46% believe in the Big Bang theory
despite all of the above being scientific facts that are undisputed in the scientific community.
There are practically zero scientists who reject evolution on scientific grounds, and those who do are often paid by religious organisations to promote their agenda i.e. they are strongly biased.
.](/post/evolution-evidence/survey_for_evo_support_huede650c85e45379e53dc3c61f7294353_224316_a064026b1ab42a05080aa09196edc51b.webp)
A significant number of scientific societies have explicitly rejected intelligent design.
Other relevant info: Level of support for evolution
Intelligent Design is Political, not Scientific
The concept of intelligent design (ID) was conceived by the Discovery Institute (DI), an evangelical Christian ’think tank’, as an attempt to circumvent the 1987 Edwards v. Aguillard court ruling prohibiting the teaching of creationism in public schools due to violation of church-state separation. ID was also intended to be more appealing to the general public, a necessary part of the DI’s “Wedge Strategy”, whose long-term goal is to effectively work towards installing theocracy in the US, as outlined in their leaked Wedge Document in 2001. Once this document was exposed, the DI published The Wedge Document: So What? in an attempt at damage control, where they both denied and doubled down on some their intentions.
As part of this effort to promote ID, the DI released a petition called “Dissent from Darwinism”, which asked a wide variety of experts including scientists, doctors and engineers whether they agreed with the following statement:
“I am skeptical of claims for the ability of random mutation and natural selection to account for the complexity of life”.
This petition was intentionally and deceptively worded, since a reasonable scientist may well agree that mutation and selection are not sufficient for evolution, since there are in fact other mechanisms of evolution (e.g. genetic drift, gene flow, sexual selection…). Additionally, by including non-biologists such as engineers and doctors, the DI deliberately inflated the number of qualified signatories, and in fact the vast majority of the signatories were not biologists, totalling about 1,000 individuals in total.
The National Center for Science Education (NCSE) put out a counter-petition called “Project Steve”, which asked scientists (not engineers or doctors) a far more precise statement to the contrary, and only signatories named “Steve” (or some close variation thereof) were counted. About 1,500 Steves signed this petition, outnumbering the original.
The teaching of ID in US public schools was challenged in the 2005 Kitzmiller v. Dover case, which ruled that ID is not science, but creationism, and therefore cannot be taught. Overwhelming evidence was brought against the ID proponents, including several refutations to talking points which are still parroted to this day by their followers. After the trial, it was noticed that the DI’s ‘creation science’ textbook Of Pandas and People had simply copy-pasted the word “design proponents” in place of “creationists” in the text, with one edition of the book featuring the telling typo, “cdesign proponentsists”.
.](/post/evolution-evidence/id_not_science_huc0eb3abbb91b89d80e7616f00aab4297_864129_2d874658e19b2d9f2577ca6a581e6353.webp)
In the run-up to the 2024 US presidential election, the DI quietly became a ‘coalition partner’ for Project 2025. Once the public became aware of Project 2025 and its draconian ambitions, the DI allegedly requested its logo be removed from the Project 2025 website to cover its tracks, but this was caught and the DI remains a coalition partner as of 2025.
.](/post/evolution-evidence/di_on_project_2025_hu7d0bea67808077ae1bfb8bec976d955c_297121_6d67ba456131fc939a4cbefbe9703bdd.webp)
Creationism is not science, and intelligent design (ID) is merely creationism with a science-coloured coat of paint. Neither of these ideas make any testable falsifiable predictions, except for those which have already been tested and falsified.
Video playlist extensively exposing the DI: here.
The DI was originally and continues to be funded by rich Christian nationalist organisations, having raked in over $15 million per year in donations as of 2021 (source: here).
Incompetence of “Creation Scientists”
“Creation scientists” are degree-holding scientists who intend to use science to find evidence for their religious convictions. Despite being qualified (sometimes), they have a reputation for being exceptionally dishonest, untrustworthy, and most importantly, highly incompetent in their work promoting creationism. Examples include:
Jeffrey Tomkins, who made numerous basic arithmetic and methodological errors in attempting to show that humans and chimpanzees share much less DNA than the conventional figure of >95%.
“Mendel’s Accountant”, a computer program written by John Sanford, intended to show that mutation would always lead to ‘genetic entropy’ (loss of genetic information and degradation). The code was found to be heavily skewed to favour this conclusion, by biassing the effect of harmful mutations, among several other flaws.
“Biblical Radiocarbon”, a website published by creationists at Answers in Genesis (AiG) running a program that aims to ‘recalibrate’ conventional radiocarbon dates into a young-earth timeline. The program’s code was found to be of exceptionally poor quality with numerous bugs.
Salvador (Sal) Cordova, a creation scientist who gave a presentation at a conventional evolution conference. The presentation was highly unprofessional and he made no attempt to communicate any of his points clearly. Dr Dan Stern Cardinale and Dr Zach Hancock reviewed the presentation here. He has also written a very poor quality paper here which was rejected from the BioArxiv preprint server as well as several journals over a course of 8 years.
Nathaniel Jeanson is a Harvard PhD who has readily admitted that he only acquired his prestigious degree in order to promote creationism with authority. He has been caught recycling arguments without addressing the counters many times.
“Raw Matt” (Matthew Nailor) is an accomplice on the ‘Standing for Truth’ YouTube channel, a bottom-tier platform for YEC apologetics. He wrote a “paper” here with some of the most terrible formatting and content imaginable, and also fraudulently copied-and-pasted the PLOS One logo onto his paper, to pretend that the PLOS journal published his paper.
It is often difficult to discern whether these “creation scientists” are incompetent or intentionally dishonest - the former seems far less likely in the case of the more qualified individuals.
There is also a group of intelligent design (ID) proponents working at the Discovery Institute (DI). Unlike the YECs, the DI’s ID proponents are all paid to lie for a specific wider agenda and can therefore only be concluded to be intentionally deceptive. They have been exposed many times, such as in this video series. Some examples:
Casey Luskin: deceptively edited a segment of a PBS Nova documentary to remove a voiceover explaining how a scientist made a plaster cast of the ‘Lucy’ (Australopithecus afarensis) pelvis fossil. He substituted this with a narrative claiming the scientists fraudulently cut and deformed the original specimen to make it falsely appear bipedal.
Günter Bechly worked at the DI and allegedly ended his life in a murder-suicide car crash, killing one other person. The DI made no mention of these allegations in their post reminiscing on his career. Sources: here and here.
Michael Behe: selectively deleted parts of a data table showing the effect of mutations in a population of polar bears to falsely claim that neutral (benign) mutations are far rarer than they actually are.
Jonathan McLatchie: taught at a Bible college (Sattler college), graduated with a PhD dissertation that is unavailable online (here, extremely unusual, potentially indicative of poor quality) and pretends to be a scientist without having done any legitimate work (his only published paper is in the DI’s in-house journal ‘Bio-Complexity’). He was exposed as incompetent in an exchange with PZ Meyers (here), and ragequit a ‘Christianity vs atheism’ debate against Matt Dillahunty (here).
Philosophy of Science
- Science rejection is linked to unjustified overconfidence in knowledge (Fonseca et al., 2023)
- Science rejection is correlated with religious intolerance (Ding, Johar & Morris, 2024)
- Evolution rejection is correlated with not understanding how science operates (Lombrozo, Thanukos & Weisberg, 2008)
- Fundamentalist religious belief is similar to belief in patients with brain lesions (Zhong et al., 2017):
[W]e found that participants with dorsolateral prefrontal cortex (dlPFC) lesions have fundamentalist beliefs similar to patients with vmPFC lesions and that the effect of a dlPFC lesion on fundamentalism was significantly mediated by decreased cognitive flexibility and openness.
Some Relevant Quotes
Nothing in biology makes sense except in the light of evolution.
— Theodosius Dobzhansky, Evolutionary Biologist
I write about biology from the point of view of a physicist. Some physicists are arrogant and some are humble. I prefer to be humble. Arrogant physicists say that biology needs better concepts; since physicists are good at concepts, our job is to tell biologists how to think. Humble physicists say that biology needs better hardware; since physicists are good at hardware, our job is to invent new tools for biologists to use. With the exception of Max Delbruck and Francis Crick and a few other pioneers in the heroic age of molecular biology, physicists who tried to teach biologists how to think have failed dismally.
— Freeman Dyson, Theoretical Physicist, as cited by Dr David Tong in his Cambridge Mathematical Biology lecture notes.
It doesn’t matter how beautiful your theory is, it doesn’t matter how smart you are. If it doesn’t agree with experiment, it’s wrong.
— Richard Feynman, Theoretical Physicist
All models are wrong, but some are useful.
— George Box, Statistician
I believe God did intend, in giving us intelligence, to give us the opportunity to investigate and appreciate the wonders of His creation. He is not threatened by our scientific adventures.
— Francis Collins, Director of the Human Genome Project and the NIH